

Scientia Silvae Sinicae ›› 2022, Vol. 58 ›› Issue (4): 74-81.doi: 10.11707/j.1001-7488.20220408
• Research papers • Previous Articles Next Articles
Wenting Pan,Jianjun Sun,Qinqin Yuan,Lili Zhang,Kangqiao Deng,Yueqiao Li*
Received:2021-04-16
Online:2022-04-25
Published:2022-07-20
Contact:
Yueqiao Li
CLC Number:
Wenting Pan,Jianjun Sun,Qinqin Yuan,Lili Zhang,Kangqiao Deng,Yueqiao Li. Analysis of Genetic Diversity and Structure in Different Provenances of Liriodendron by RAD-seq Technique[J]. Scientia Silvae Sinicae, 2022, 58(4): 74-81.
Table 1
Geographic information and samplenumber of 13 provenances in the experiment"
| 种源 Provenance | 种源地 Provenance region | 北纬 Latitude | 东经 Longitude | 海拔 Elevation/m | 样品数 Number of samples |
| XY | 四川叙永 Sichuang Xuyong | 28.20°N | 105.5°E | 700~900 | 10 |
| EX | 湖北鄂西 Hubei Exi | 30.30°N | 109.0°E | 1 100~1 300 | 11 |
| MN | 贵州陌南 Guizhou Monan | 26.80°N | 109.5°E | 950~1 150 | 11 |
| LY | 湖南浏阳 Hunan Liuyang | 28.05°N | 113.9°E | 1 000~1 200 | 11 |
| SZ | 湖南桑植 Hunan Sangzhi | 29.15°N | 110.2°E | 1 100~1 300 | 11 |
| LS | 江西庐山 Jiangxi Lushan | 29.53°N | 116.0°E | 1 000~1 200 | 10 |
| WYS | 江西武夷山 Jiangxi Wuyishan | 27.92°N | 117.8°E | 900~1 100 | 10 |
| MSL | 密苏里州 Missouri | 37.60°N | 91.5°W | 600~800 | 11 |
| LYS | 路易斯安纳州 Louisiana | 31.00°N | 93.6°W | 0~100 | 10 |
| BK | 北卡罗来纳州 North Carolina | 35.13°N | 80.1°W | 1 000~1 200 | 12 |
| NK | 南卡罗来纳州 South Carolina | 34.50°N | 81.8°W | 100~300 | 14 |
| YN | 云南勐腊 Yunnan Mengla | 21.40°N | 101.6°E | 600~800 | 11 |
| DBS | 大别山舒城 Dabieshan Shucheng | 31.13°N | 118.2°E | 1 350~1 550 | 11 |
Table 2
Statistics of RAD-seq sequencing data of 13 provenances"
| 种源 Provenance | 总碱基数 Total nucleotides (Gb) | 有效数据 Clean reads (Mb) | RAD标记数 RAD tag | 测序深度 Sequencing depth (x) |
| XY | 1.77 | 5.90 | 129 420 | 16.29 |
| EX | 2.46 | 8.19 | 140 080 | 20.19 |
| MN | 4.20 | 14.01 | 155 030 | 34.20 |
| LY | 1.94 | 6.46 | 147 270 | 17.77 |
| SZ | 1.41 | 4.70 | 157 950 | 13.70 |
| LS | 4.37 | 14.57 | 156 650 | 35.74 |
| WYS | 2.71 | 9.02 | 128 700 | 25.34 |
| MSL | 3.89 | 12.97 | 136 120 | 39.76 |
| LYS | 3.75 | 12.50 | 139 340 | 34.95 |
| BK | 1.99 | 6.62 | 140 020 | 19.14 |
| NK | 3.14 | 10.48 | 128 720 | 29.78 |
| YN | 2.15 | 7.16 | 127 330 | 19.47 |
| DBS | 2.16 | 7.20 | 149 230 | 18.97 |
| 均值Mean | 2.76 | 9.20 | 142 130 | 24.76 |
Table 3
Statistics of genetic structure in different provenances"
| 种源 Provenance | 等位基因数 Allele number | Ho Observed heterozygosity | He Expected heterozygosity | π Nucleotide diversity | Gst Gene differential coefficient | Nm Gene flow |
| XY | 4 334 | 0.035 2 | 0.059 7 | 0.064 0 | 0.211 2 | 0.933 8 |
| EX | 5 573 | 0.049 6 | 0.058 1 | 0.061 7 | 0.196 8 | 1.020 3 |
| MN | 3 834 | 0.039 5 | 0.047 2 | 0.049 9 | 0.243 0 | 0.778 8 |
| LY | 2 934 | 0.036 4 | 0.049 9 | 0.053 0 | 0.237 8 | 0.801 3 |
| SZ | 4 212 | 0.036 9 | 0.052 0 | 0.055 5 | 0.213 1 | 0.923 4 |
| LS | 855 | 0.044 8 | 0.045 9 | 0.048 9 | 0.235 4 | 0.812 2 |
| WYS | 758 | 0.038 9 | 0.044 8 | 0.047 3 | 0.234 9 | 0.814 4 |
| MSL | 1 113 | 0.038 6 | 0.038 4 | 0.040 6 | 0.439 8 | 0.318 4 |
| LYS | 2 726 | 0.043 8 | 0.055 1 | 0.058 8 | 0.385 5 | 0.398 5 |
| BK | 2 017 | 0.044 2 | 0.058 1 | 0.061 6 | 0.367 9 | 0.429 5 |
| NK | 3 309 | 0.047 1 | 0.058 8 | 0.061 6 | 0.361 5 | 0.441 6 |
| YN | 2 604 | 0.013 2 | 0.029 9 | 0.031 6 | 0.280 2 | 0.642 2 |
| DBS | 1 245 | 0.024 4 | 0.022 2 | 0.023 6 | 0.324 8 | 0.519 8 |
| 均值 Mean | 2 732 | 0.037 9 | 0.047 7 | 0.050 6 | 0.287 1 | 0.679 6 |
Table 4
Subordinative function values of genetic diversity parameters among 13 provenances in Liriodendron"
| 参数 parameters | XY | EX | MN | LY | SZ | LS | WYS | MSL | LYS | BK | NK | YN | DBS |
| Ho得分Ho score | 0.60 | 1.00 | 0.72 | 0.64 | 0.65 | 0.87 | 0.71 | 0.70 | 0.84 | 0.85 | 0.93 | 0.00 | 0.31 |
| He得分He score | 1.00 | 0.96 | 0.67 | 0.74 | 0.79 | 0.63 | 0.60 | 0.43 | 0.88 | 0.96 | 0.98 | 0.21 | 0.00 |
| π得分π score | 1.00 | 0.94 | 0.65 | 0.73 | 0.79 | 0.63 | 0.59 | 0.42 | 0.87 | 0.94 | 0.94 | 0.20 | 0.00 |
| 综合得分 Comprehensive score | 2.60 | 2.90 | 2.04 | 2.10 | 2.24 | 2.13 | 1.90 | 1.55 | 2.59 | 2.75 | 2.85 | 0.40 | 0.31 |
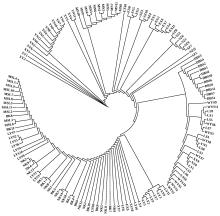
Fig.2
Neighbour-joining phylogram illustrating genetic relationships among 143 individuals Each branch represents one piece of germplasm, and the length of the branch represents the evolutionary distance between two species, the number on a branch of the tree represents the percentage of support for that branch."
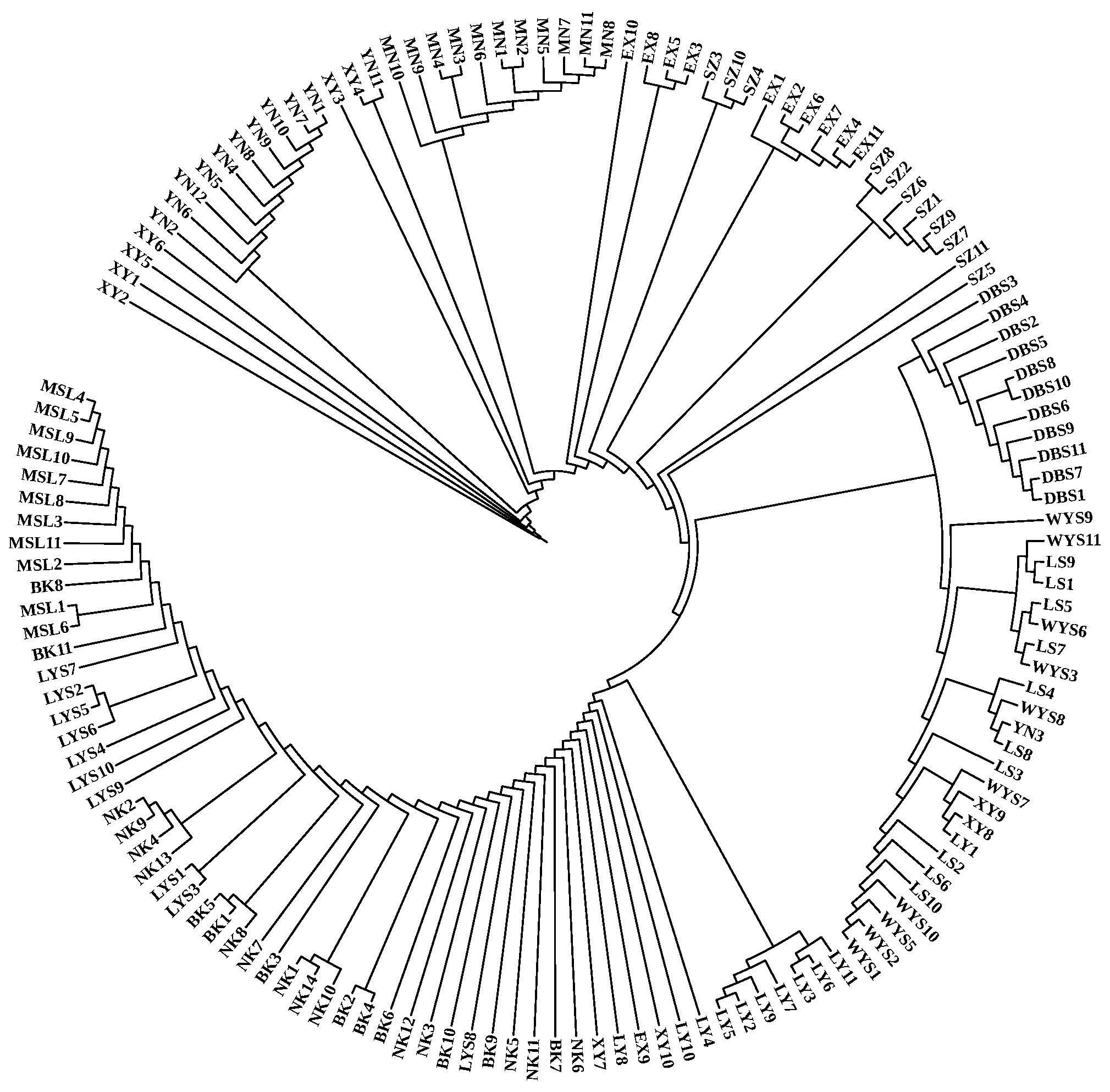
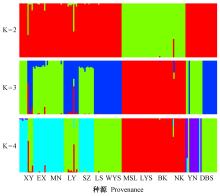
Fig.3
Bar plot of 2-4 clusters identified with adegenet R package The picture left shows the clustering of 143 samples with cluster values from 2 to 4 (each color represents a subgroup, Each column represents a sample). The figure on the right shows the CV error value corresponding to each K value (When K is 3, the CV error value is minimal)."
| 郝日明, 贺善安, 汤诗杰, 等. 鹅掌楸在中国的自然分布及其特点. 植物资源与环境, 1995, 4 (1): 1- 6. | |
| Hao R M , He S A , Tang S J , et al. Geographical distribution of Liriodendron chinense in China and its significance. Plant Resources and Environment, 1995, 4 (1): 1- 6. | |
| 黄承玲, 姚刚, 田晓玲, 等. 基于RAD高通量测序的贵州百里杜鹃保护区杜鹃花属分类. 林业科学, 2021, 57 (2): 72- 81. | |
| Huang C L , Yao G , Tian X L , et al. Phylogenomic analysis of rhododendron species in Guizhou Baili rhododendron reserve based on RAD sequencing. Scientia Silvae Sinicae, 2021, 57 (2): 72- 81. | |
| 李康琴. 2013. 鹅掌楸属群体遗传结构及分子系统地理学研究. 南京: 南京林业大学. | |
| Li K Q. 2013. Studies on population genetics and molecular phylogeography of Liriodendron. Nanjing: Nanjing Forestry University. [in Chinese] | |
| 李云飞, 李世明, 金鑫, 等. 基于RAD高通量测序探讨中国85种杜鹃花属植物的分类. 林业科学研究, 2019, 32 (3): 1- 8. | |
| Li Y F , Li S M , Jin X , et al. Phylogenomic analysis of 85 rhododendron species in China based on RAD sequencing. Forest Research, 2019, 32 (3): 1- 8. | |
| 陆叶, 龙晓飞, 王鹏凯, 等. 基于RAD-seq技术的鹅掌楸基因组SNP标记开发. 南京林业大学学报(自然科学版), 2019, 43 (4): 1- 7. | |
| Lu Y , Long X F , Wang P K , et al. Development of genomic SNP markers based on RAD-seq and genome data in Liriodendron. Journal of Nanjing Forestry University (Natural Sciences Edition), 2019, 43 (4): 1- 7. | |
| 罗群凤, 胥猛, 冯源恒, 等. 北美鹅掌楸LtCHS基因的克隆及生物信息学与组织表达特征分析. 林业科学, 2015, 51 (5): 37- 45. | |
| Luo Q F , Xu M , Feng Y H , et al. Cloning and bioinformatics of chalcone synthase gene (CHS) in Liriodendron tulipifera and characterization of its tissue expression. Scientia Silvae Sinicae, 2015, 51 (5): 37- 45. | |
| 彭松, 马淼, 郑勇奇, 等. 不同种源花楸树幼苗越夏能力的比较. 生态学杂志, 2014, 33 (2): 321- 327. | |
| Peng S , Ma M , Zheng Y Q , et al. Comparison in thermotolerance over summer of seedlings among different provenances of Sorbus pohuashanensis (Hance) Hedl. Chinese Journal of Ecology, 2014, 33 (2): 321- 327. | |
| 祁荔. 2017. 鹅掌楸遗传图谱构建及重要性状QTL初步定位. 南京: 南京林业大学. | |
| Qi L. 2017. Genetic linkage map construction and QTL mapping of Liriodendron. Nanjing: Nanjing Forestry University. [in Chinese] | |
| 王晓阳, 李火根. 鹅掌楸苗期生长杂种优势的SSR分析. 林业科学, 2011, 47 (4): 57- 62. | |
| Wang X Y , Li H G . Possible mechanism analysis for heterosis of hybrid Liriodendron based on seedling growth and SSR markers. Scientia Silvae Sinicae, 2011, 47 (4): 57- 62. | |
| 王章荣. 鹅掌楸属树种杂交育种与利用. 北京: 中国林业出版社, 2005. | |
| Wang Z R . With the use of Liriodendron hybrids breeding. Beijing: China Forestry Publishing House, 2005. | |
| 赵亚琦, 成铁龙, 施季森, 等. 鹅掌楸属SRAP分子标记体系优化及遗传多样性分析. 林业科学, 2014, 50 (7): 37- 43. | |
| Zhao Y Q , Cheng T L , Shi J S , et al. Optimization of a SRAP-PCR system for analysis of genetic diversity of Liriodendron. Scientia Silvae Sinicae, 2014, 50 (7): 37- 43. | |
| 朱晓琴, 马建霞, 姚青菊, 等. 鹅掌楸遗传多样性的等位酶论证. 植物资源与环境, 1995, 4 (3): 9- 14. | |
| Zhu X Q , Ma J X , Yao Q J , et al. Allozyme verification on the population of Liriodendron chinense. Plant Resources and Environment, 1995, 4 (3): 9- 14. | |
|
Catchen J , Hohenlohe P , Bassham S , et al. Stacks: an analysis tool set for population genomics. Molecular Ecology, 2013, 22 (11): 3124- 3140.
doi: 10.1111/mec.12354 |
|
|
Feng J Y , Zhao S , Li M , et al. Genome-wide genetic diversity detection and population structure analysis in sweet potato (Ipomoea batatas) using RAD-seq. Genomics, 2020, 112 (2): 1978- 1987.
doi: 10.1016/j.ygeno.2019.11.010 |
|
| Jérémy G , Charlotte M , Tomasz S , et al. Disco Snp-RAD: de novo detection of small variants for RAD-Seq population genomics. PeerJ, 2020, 8 (2): e9291. | |
|
Lexander D H , Novembre J , Lange K . Fast model-based estimation of ancestry in unrelated individual. Genome Research, 2009, 19 (9): 1655- 1664.
doi: 10.1101/gr.094052.109 |
|
|
Li Z B , Hua Z T , Dong L , et al. Quantitative trait locus mapping for yield-associated agronomic traits in a BC2F6 population of Japonica hybrid rice Liaoyou 5218. Journal of Plant Growth Regulation, 2020, 39 (1): 60- 71.
doi: 10.1007/s00344-019-09963-4 |
|
|
Liu L W , Zhao L P , Gong Y Q , et al. DNA fingerprinting and genetic diversity analysis of late-bolting radish cultivars with RAPD, ISSR and SRAP markers. Scientia Horticulturae, 2008, 116 (3): 240- 247.
doi: 10.1016/j.scienta.2007.12.011 |
|
|
Miller M R , Dunham J P , Amores A , et al. Rapid and cost-effective polymorphism identification and genotyping using restriction site associated DNA (RAD) markers. Genome Research, 2007, 17 (2): 240- 248.
doi: 10.1101/gr.5681207 |
|
|
Parks C R , Wendel J F . Molecular divergence between Asian and North American species of Liriodendron (Magnoliaceae) with implications for interpretation of fossil floras. American Journal of Botany, 1990, 77 (10): 1243- 1256.
doi: 10.1002/j.1537-2197.1990.tb11376.x |
|
|
Peterson B K , Weber J N , Kay E H , et al. Double digest RADseq: an inexpensive method for de novo SNP discovery and genotyping in model and non-model species. PLoS One, 2012, 7 (5): e37135.
doi: 10.1371/journal.pone.0037135 |
|
|
Sheng Y , Hao Z D , Peng Y , et al. Morphological, phenological, and transcriptional analyses provide insight into the diverse flowering traits of a mutant of the relic woody plant Liriodendron chinense. Horticulture Research, 2021, 8 (1): 174.
doi: 10.1038/s41438-021-00610-2 |
|
|
Tamura K , Peterson D , Peterson N , et al. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Molecular Biology and Evolution, 2011, 28 (10): 2731- 2739.
doi: 10.1093/molbev/msr121 |
|
| Wang Y , Shahid M Q , Lin S , et al. Footprints of domestication revealed by RAD-tag resequencing in loquat: SNP data reveals a non-significant domestication bottle neck and a single domestication event. BMC Genomics, 2017, 18 (1): 354. | |
| Yeh F C, Yang R C. 2000. Popgene version 1. 32. http://www.ualberta.ca/~fyeh/download.htm. | |
| Zaouali Y , Chograni H , Trimech R , et al. Genetic diversity and population structure among Rosmarinus officinalis L.(Lamiaceae) varieties: var. typicus Batt. and var. troglodytorum Maire. based on multiple traits. Industrial Crops and Products, 2012, 38, 166- 176. |
| [1] | Chaoqun Du,Xiaomei Sun,Yunhui Xie,Yimei Hou. Genetic Diversity of Larix kaempferi Populations with Different Levels of Improvement in Northern Subtropical Region [J]. Scientia Silvae Sinicae, 2021, 57(5): 68-76. |
| [2] | Chengling Huang,Gang Yao,Xiaoling Tian,Yongquan Ren,Jiayong Huang,Yongpeng Ma. Phylogenomic Analysis of Rhododendron Species in Guizhou Baili Rhododendron Reserve Based on RAD Sequencing [J]. Scientia Silvae Sinicae, 2021, 57(2): 72-81. |
| [3] | Huanhuan Chen,Gexi Xu,Fanqiang Ma,Shun Liu,Miaomiao Zhang,Xiangwen Cao,Jian Chen,Guangdong Zhao,Hongguo Yang,Zuomin Shi. Phylogenetic Structure of Undergrowth Layers across Subalpine Dark Coniferous Forests and Their Post-Harvesting Secondary Forests in Western Sichuan [J]. Scientia Silvae Sinicae, 2020, 56(7): 1-11. |
| [4] | Yihong Meng,Gangbiao Xu,Mengzhu Lu,Xiaolong Jiang,Feilong Guo. Population Genetic Structure and Demographic History of Disanthus cercidifolius var. longipes [J]. Scientia Silvae Sinicae, 2020, 56(7): 55-62. |
| [5] | Xia Li,Libao Wang,Yafeng Wen,Jun Lin,Xingtong Wu,Meiling Yuan,Yuan Zhang,Minqiu Wang,Xinyu Li. Genetic Diversity of Chinese Fir (Cunninghamia lanceolata) Breeding Populations among Different Generations [J]. Scientia Silvae Sinicae, 2020, 56(11): 53-61. |
| [6] | Ma Songmei, Wang Chuncheng, Sun Fangfang, Wei Bo, Nie Yingbin. Genetic Diversity of an Endangered Plant Amygdalus ledebouriana in Xinjiang [J]. Scientia Silvae Sinicae, 2019, 55(9): 71-80. |
| [7] | Zhang Shouke, Fang Linxin, Liu Yaning, Wang Yi, Zhang Wei, Shu Jinping, Zhang Yabo, Wang Yangdong, Wang Haojie. Genetic Differentiation and Structural Variation of ATP Synthase Gene of Curculio chinensis (Coleptera: Curculionidae) under Selection Pressure at Different Altitudes [J]. Scientia Silvae Sinicae, 2019, 55(6): 65-73. |
| [8] | Chen Tingting, Wang Pengkai, Zhang Jiaji, Shi Jisen, Cheng Tielong, Chen Jinhui. Effects of Combined ABA and ZT Treatment on Somatic Embryogenesis and Development of Liriodendron sino-americanum [J]. Scientia Silvae Sinicae, 2019, 55(3): 64-71. |
| [9] | Jin Ling, Liu Mingguo, Dong Shengjun, Wu Yueliang, Zhang Xin. Genetic Diversity and Fingerprints of 97 Armeniaca sibirica Clones Based on SSR Markers [J]. Scientia Silvae Sinicae, 2018, 54(7): 51-61. |
| [10] | Hao Minhui, Li Xiaoyu, Xia Mengjie, He Huaijiang, Zhang Chunyu, Zhao Xiuhai. Effects of Tending Felling on Functional and Phylogenetic Structures in a Multi-Species Temperate Secondary Forest at Jiaohe in Jilin Province [J]. Scientia Silvae Sinicae, 2018, 54(5): 1-9. |
| [11] | Bao Wenquan, Wuyun Tana, Du Hongyan, Li Tiezhu, Liu Huimin, Wang Lin, Bai Yu. Genetic Diversity and Population Structure of Amygdalus mira in the Tibet Plateau in China Based on SSR Markers [J]. Scientia Silvae Sinicae, 2018, 54(2): 30-41. |
| [12] | Yang Hanbo, Zhang Rui, Wang Bangshun, Xu Zhaoyou, Chen Huanwei, Zhou Zhichun. Analysis of Genetic Diversity in Schima superba Plus Tree Germplasms by SSR Markers [J]. Scientia Silvae Sinicae, 2017, 53(5): 43-53. |
| [13] | Mao Xiuhong, Zheng Yongqi, Sun Baiyou, Zhang Yuanshuai, Han Congcong, Wei Xiao, Xun Shouhua. Genetic Diversity and Fingerprints of Robinia pseudoacacia Clones Based on SSR Markers [J]. Scientia Silvae Sinicae, 2017, 53(10): 80-89. |
| [14] | Feng Yuanheng, Li Huogen, Yang Zhangqi, Huang Yongli, Luo Qunfeng, Zhang Yuan. Construction of Second Generation Breeding Population of Pinus massoniana in Guangxi [J]. Scientia Silvae Sinicae, 2017, 53(1): 54-61. |
| [15] | Song Zhijiao, Yang Heyu, Weng Qijie, Zhou Changpin, Li Fagen, Li Mei, Lu Wanhong, Luo Jianzhong, Gan Siming. Genetic Diversity and Selective Loci in Eucalyptus tereticornis Populations [J]. Scientia Silvae Sinicae, 2016, 52(9): 39-47. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||