

林业科学 ›› 2026, Vol. 62 ›› Issue (5): 1-15.doi: 10.11707/j.1001-7488.LYKX20260014
收稿日期:2026-01-08
修回日期:2026-03-15
出版日期:2026-05-10
发布日期:2026-05-12
通讯作者:
赵振利
E-mail:zlxx64@126.com
基金资助:
Ruoyu Wang,Mengqi Feng,Lu Sima,Zhenli Zhao*( ),Guoqiang Fan
),Guoqiang Fan
Received:2026-01-08
Revised:2026-03-15
Online:2026-05-10
Published:2026-05-12
Contact:
Zhenli Zhao
E-mail:zlxx64@126.com
摘要:
目的: 为解析白花泡桐木质素生物合成途径中关键限速酶肉桂酰辅酶A还原酶(CCR)的基因组学特征及功能分化,基于白花泡桐全基因组数据,对CCR家族进行全基因组鉴定和系统分析,探究白花泡桐在木质素合成及生物和非生物胁迫下的响应机制,为基因编辑技术定向培育优良抗逆泡桐新品种提供理论参考。方法: 以拟南芥中已知的CCR蛋白序列为诱饵,基于泡桐基因组数据库进行BLAST比对,获得白花泡桐PfCCRs家族成员,并利用生物信息学技术分析其家族成员进化关系、启动子结合元件、共线性等特征,采用实时荧光定量PCR(RT-qPCR)技术对白花泡桐PfCCRs在不同组织以及逆境胁迫下的表达水平进行检测。结果: 1) 在白花泡桐全基因组中鉴定到14个PfCCRs成员,氨基酸长度111~360 aa,分为Cluster Ⅰ~Cluster Ⅴ 5个亚家族,均含有NADB_Rossmann结构域。2) PfCCRs启动子区域富集光响应、激素信号响应和逆境胁迫响应相关元件,且PfCCRs不均匀分布于8条染色体,其分别与双子叶植物芝麻和拟南芥CCRs之间产生10个共线基因对。3) 系统发育树构建与分析表明,PfCCR9和PfCCR11与拟南芥木质素合成调控因子AtCCR1以及抗病防御相关基因AtCCR2共同聚集于Cluster Ⅰ,此外,在干旱、盐胁迫下PfCCR9和PfCCR11的表达显著上调,表明其在抗逆分子调控中发挥关键作用;而PfCCR12在丛枝植原体(PaWB)感染苗中的表达显著上调,表明其在病原体胁迫中起重要功能。4) PfCCR1、PfCCR3、PfCCR6、PfCCR7、PfCCR11、PfCCR13和PfCCR14在茎中显著高表达,而PfCCR10和PfCCR11在根中显著高表达,表明这8个基因在木质素合成中发挥重要作用。结论: 本研究鉴定调控泡桐木质素合成、抗逆等分子机制中发挥作用的PfCCRs家族关键成员,根据进化分析与表达量验证预测Cluster Ⅰ中的PfCCR9和PfCCR11以及Cluster Ⅰ中PfCCR12的在抗逆胁迫中具有重要调控作用,为泡桐抗逆新品种选育的候选基因。
中图分类号:
王若雨,冯梦琦,司马璐,赵振利,范国强. 白花泡桐PfCCR全基因组鉴定及逆境响应[J]. 林业科学, 2026, 62(5): 1-15.
Ruoyu Wang,Mengqi Feng,Lu Sima,Zhenli Zhao,Guoqiang Fan. Genome-wide Identification and Stress Response of PfCCR Gene Family in Paulownia fortunei[J]. Scientia Silvae Sinicae, 2026, 62(5): 1-15.
表1
引物序列列表"
| 引物名称 Primer names | 正向引物序列(5'?3') Forward primer sequences (5'?3') | 反向引物序列(5'?3') Reverse primer sequences (5'?3') |
| PfCCR1 | ACCACGTCAACCTCCTCAAC | CGGCAAGCACGTTCAGAGTA |
| PfCCR2 | AACGTGCTTGGATCTTGTGC | ACCGCTTCAGATGTTCGAGG |
| PfCCR3 | ATTGAGGAGACTTGGTGGGC | TTGTGTTGAGTGTTGGCTGC |
| PfCCR4 | ATGAGACTTGGTGGGCTGAC | CTTGTGTTGAGCGTTGGCTG |
| PfCCR5 | ATTGACGGCTGTGATGGTGT | ACTGAAGCCACCGATGATGT |
| PfCCR6 | TGGTGAATCCTGGAACCGTG | CACAAGTGCCTTCCAGTTGC |
| PfCCR7 | TGGTGAATCCAGGAACGGTG | TCCAGTGGCCGATGTGTTTT |
| PfCCR8 | TGTGGACCCTGCTGGACTAT | CACTCCAGCACTTGTGGTCA |
| PfCCR9 | ACGGTTGCGATGGTGTTTTC | CACGGCTGACCTTAGCTTCT |
| PfCCR10 | TGTTCTTGGGTCATGTGCGA | CGGACCACCAAGTCTCATCA |
| PfCCR11 | TCAGCCAACTGTGAATGCCA | GCCAATGCCACATCCTTGAC |
| PfCCR12 | ATGTTCATGGCACCGTCAGA | CACCTCCGGGTTTGGTACAG |
| PfCCR13 | GTGCTTGTGGCCTACGATGA | GGGGACACTTCTTCCACGTC |
| PfCCR14 | AGTTGTCCACCTAAAGGCACT | AGTGTTCCTTTCACTGCTGGT |
| Pfactin | AATGGAATCTGCTGGAAT | ACTGAGGACAATGTTACC |
表2
PfCCRs的理化性质分析"
| 基因名称 Gene name | 基因ID Gene ID | 氨基酸数目 Amino acid/aa | 分子质量 Molecular mass/Da | 等电点 pI | 不稳定指数 Instability index | 亲水性均值 Hydrophilic mean value | 亚细胞定位 Subcellular localization |
| Pfo02g012130.1 | PfCCR1 | 339 | 6.14 | 31.26 | ?0.003 | 细胞质、叶绿体Cytoplasm, chloroplast | |
| Pfo05g006750.1 | PfCCR2 | 323 | 7.50 | 22.25 | ?0.135 | 叶绿体Chloroplast | |
| Pfo05g006760.1 | PfCCR3 | 323 | 6.99 | 24.66 | ?0.120 | 叶绿体Chloroplast | |
| Pfo05g006770.1 | PfCCR4 | 323 | 6.20 | 22.45 | ?0.122 | 叶绿体Chloroplast | |
| Pfo05g006780.1 | PfCCR5 | 324 | 5.76 | 30.56 | ?0.003 | 叶绿体Chloroplast | |
| Pfo05g013040.1 | PfCCR6 | 334 | 6.12 | 38.23 | ?0.078 | 叶绿体、细胞质Chloroplast, cytoplasm | |
| Pfo05g013050.1 | PfCCR7 | 210 | 5.76 | 38.83 | 0.023 | 叶绿体Chloroplast | |
| Pfo06g015420.1 | PfCCR8 | 360 | 5.90 | 37.74 | ?0.090 | 叶绿体、细胞质、细胞核 Chloroplast, cytoplasm, nucleus | |
| Pfo08g010280.1 | PfCCR9 | 332 | 6.52 | 27.42 | ?0.145 | 叶绿体Chloroplast | |
| Pfo12g004120.1 | PfCCR10 | 324 | 7.04 | 26.36 | ?0.040 | 叶绿体Chloroplast | |
| Pfo14g003150.1 | PfCCR11 | 332 | 6.03 | 30.86 | ?0.172 | 叶绿体Chloroplast | |
| Pfo15g012660.1 | PfCCR12 | 347 | 6.67 | 27.05 | ?0.148 | 叶绿体Chloroplast | |
| Pfo17g001580.1 | PfCCR13 | 327 | 5.22 | 43.36 | ?0.175 | 细胞质、细胞核Cytoplasm, nucleus | |
| Pfo17g009820.1 | PfCCR14 | 111 | 6.90 | 46.50 | 0.195 | 叶绿体、细胞质Chloroplast, cytoplasm |
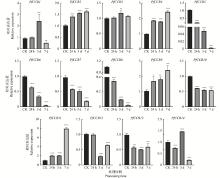
图7
干旱胁迫下PfCCRs基因表达模式分析 采用RT-qPCR方法分析PfCCRs在干旱胁迫下的表达模式,Pfactin作内参基因,以CK作为对照(相对表达水平设为1.0)。CK:喷施等量水作为对照;24 h:150 g·L−1 PEG 6000喷施24 h取样后为试验组;3 d:150 g·L−1 PEG 6000喷施3 天后取样为试验组;7 d:150 g·L−1 PEG 6000喷施7 天后取样为试验组。误差线表示3个生物学重复的平均标准误差。使用t-test进行差异显著性分析(*: P<0.05,**: P<0.01,***: P<0.001,****: P<0.000 1)。"
|
陈碧华. 马尾松肉桂酰辅酶A还原酶基因(CCR)克隆及分析. 林业科学, 2009, 12 (12): 46- 53.
doi: 10.3321/j.issn:1001-7488.2009.12.008 |
|
|
Chen B H. Cloning and sequence analysis of cinnamoyl-CoA reductase gene (CCR) of Pinus massoniana. Scientia Silvae Sinicae, 2009, 12 (12): 46- 53.
doi: 10.3321/j.issn:1001-7488.2009.12.008 |
|
|
董占强, 翟晓巧, 范国强, 等. 泡桐叶片RNA提取方法的研究. 河南农业大学学报, 2009, 43 (1): 40- 43.
doi: 10.16445/j.cnki.1000-2340.2009.01.016 |
|
|
Dong Z Q, Zhai X Q, Fan G Q, et al. Study on extraction methods of RNA from Paulownia leaves. Journal of Henan Agricultural University, 2009, 43 (1): 40- 43.
doi: 10.16445/j.cnki.1000-2340.2009.01.016 |
|
| 侯维海, 王建林, 旦 巴, 等. 甘蓝型和白菜型油菜肉桂酰辅酶A还原酶1基因的克隆与表达. 西北农林科技大学学报(自然科学版), 2017, 45 (11): 27- 35. | |
| Hou W H, Wang J L, Dan B, et al. Cloning and expression of cinnamoyl-CoA reductase 1 (CCR1) gene of Brassica napus L. and Brassica rapa L. Journal of Northwest A&F University (Natural Science Edition), 2017, 45 (11): 27- 35. | |
| 胡嘉祺, 齐 芪, 蒋湘宁, 等. 毛白杨融合基因4CL1-CCR对烟草木质素沉积的影响. 林业科学, 2020, 56 (10): 63- 69. | |
| Hu J Q, Qi Q, Jiang X N, et al. Effect of fusion gene 4CL1-CCR of Populus tomentosa on lignin deposition in tobacco. Scientia Silvae Sinicae, 2020, 56 (10): 63- 69. | |
| 蔺占兵, 马庆虎, 麻 密. 2001. 小麦中两个肉桂酰辅酶A还原酶基因的分离和表达分析. 植物学报, 43(10): 1043−1046. | |
| Lin Z B, Ma Q H, Mi M. Cloning and expression analysis of two wheat cDNAs encoding cinnamoyl- CoA reductase. Acta Botanica Sinica, 2001, 43(10): 1043−1046. [in Chinese] | |
|
唐雪飞, 蒲婷婷, 李豫杰, 等. 泡桐研究进展与展望. 林业科学, 2025, 61 (7): 114- 128.
doi: 10.11707/j.1001-7488.LYKX20250357 |
|
|
Tang X F, Pu T T, Li Y J, et al. Research progress and prospects of Paulownia. Scientia Silvae Sinicae, 2025, 61 (7): 114- 128.
doi: 10.11707/j.1001-7488.LYKX20250357 |
|
| 王 哲. 2020. 白花泡桐丛枝病发生相关ceRNA研究. 郑州: 河南农业大学. | |
| Wang Z. 2020. Research of ceRNA related to the occurrence of Paulownia fortunei witches’ broom. Zhengzhou: Henan Agricultural University. [in Chinese] | |
|
徐 浩, 杨克彬, 朱成磊, 等. 毛竹肉桂酰辅酶A还原酶基因PeCCR功能初步研究. 林业科学研究, 2020, 33 (2): 77- 84.
doi: 10.13275/j.cnki.lykxyj.2020.02.010 |
|
|
Xu H, Yang K B, Zhu C L, et al. Preliminary study on the function of cinnamoyl- CoA reductase gene PeCCR of moso bamboo (Phyllostachys edulis). Forest Research, 2020, 33 (2): 77- 84.
doi: 10.13275/j.cnki.lykxyj.2020.02.010 |
|
|
游 韧, 邓湘雯, 胡彦婷, 等. 树木对干旱胁迫及复水的生理生态响应研究进展. 林业科学, 2023, 59 (11): 124- 136.
doi: 10.11707/j.1001-7488.LYKX20220250 |
|
|
You R, Deng X W, Hu Y T, et al. Progress on physiological and ecological responses of trees to drought stress and rewatering. Scientia Silvae Sinicae, 2023, 59 (11): 124- 136.
doi: 10.11707/j.1001-7488.LYKX20220250 |
|
|
张嫚嫚, 刘宝光, 顾宸瑞, 等. 转BpCCR1正义链及反义链对7年生盆栽白桦木质素的影响及优良株系选择. 北京林业大学学报, 2019, 41 (6): 86- 95.
doi: 10.13332/j.1000-1522.20180412 |
|
|
Zhang M M, Liu B G, Gu C R, et al. Effects of transgenic sense and antisense of BpCCR1 on 7-year-old potted birch and selection of excellent lines. Journal of Beijing Forestry University, 2019, 41 (6): 86- 95.
doi: 10.13332/j.1000-1522.20180412 |
|
| 赵佳玥, 宗志洁, 彭玺元, 等. 2026. 真菌降解木质素的酶及其代谢途径研究进展. 林业科学, 62(3): 1−12. | |
| Zhao J Y, Zong Z J, Peng X Y, et al. 2026. An overview of novel enzymes and pathways involved in lignin biodegradation by fungi. Scientia Silvae Sinicae, 62(3): 1−12. [in Chinese] | |
|
朱 凡, 雷佳奇, 黄鑫浩, 等. 木本植物光反应对重金属胁迫响应机制的研究进展. 中南林业科技大学学报, 2022, 42 (10): 9- 21.
doi: 10.14067/j.cnki.1673-923x.2022.10.002 |
|
|
Zhu F, Lei J Q, Huang X H, et al. Advances in response mechanism of the light reaction of woody plants to heavy metal stress. Journal of Central South University of Forestry & Technology, 2022, 42 (10): 9- 21.
doi: 10.14067/j.cnki.1673-923x.2022.10.002 |
|
|
Ahuja I, Kissen R, Bones A M. Phytoalexins in defense against pathogens. Trends in Plant Science, 2012, 17 (2): 73- 90.
doi: 10.1016/j.tplants.2011.11.002 |
|
|
Bailey T L, Johnson J, Grant C E, et al. The MEME Suite. Nucleic Acids Research, 2015, 43 (W1): W39- 49.
doi: 10.1093/nar/gkv416 |
|
|
Bang S W, Choi S, Jin X, et al. Transcriptional activation of rice CINNAMOYL-CoA REDUCTASE 10 by OsNAC5, contributes to drought tolerance by modulating lignin accumulation in roots. Plant Biotechnology Journal, 2022, 20 (4): 736- 747.
doi: 10.1111/pbi.13752 |
|
|
Boerjan W, Ralph J, Baucher M. Lignin biosynthesis. Annual Review of Plant Biology, 2003, 54, 519- 546.
doi: 10.1146/annurev.arplant.54.031902.134938 |
|
|
Cao Y, Sun G, Zhai X, et al. Genomic insights into the fast growth of paulownias and the formation of Paulownia witches’ broom. Molecular Plant, 2021, 14 (10): 1668- 1682.
doi: 10.1016/j.molp.2021.06.021 |
|
| Cao Y P, Li K, Li Y L, et al. MYB transcription factors as regulators of secondary metabolism in plants. Biology, 2020, 9 (3): 61. | |
| Carocha V, Soler M, Hefer C, et al. Genome-wide analysis of the lignin toolbox of Eucalyptus grandis. New Phytologist, 2015, 206 (4): 1297- 1313. | |
|
Chabannes M, Barakate A, Lapierre C, et al. Strong decrease in lignin content without significant alteration of plant development is induced by simultaneous down-regulation of cinnamoyl CoA reductase (CCR) and cinnamyl alcohol dehydrogenase (CAD) in tobacco plants. The Plant Journal, 2001, 28 (3): 257- 270.
doi: 10.1046/j.1365-313X.2001.01140.x |
|
|
Chen C, Chen H, Zhang Y, et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Molecular Plant, 2020, 13 (8): 1194- 1202.
doi: 10.1016/j.molp.2020.06.009 |
|
|
Costa M A, Collins R E, Anterola A M, et al. An in silico assessment of gene function and organization of the phenylpropanoid pathway metabolic networks in Arabidopsis thaliana and limitations thereof. Phytochemistry, 2003, 64 (6): 1097- 1112.
doi: 10.1016/S0031-9422(03)00517-X |
|
|
De Meester B, de Vries L, Özparpucu M, et al. Vessel-specific reintroduction of CINNAMOYL-COA REDUCTASE1 (CCR1) in dwarfed ccr1 mutants restores vessel and xylary fiber integrity and increases biomass. Plant Physiology, 2018, 176 (1): 611- 633.
doi: 10.1104/pp.17.01462 |
|
|
Fan L, Linker R, Gepstein S, et al. Progressive inhibition by water deficit of cell wall extensibility and growth along the elongation zone of maize roots is related to increased lignin metabolism and progressive stelar accumulation of wall phenolics. Plant Physiology, 2006, 140 (2): 603- 612.
doi: 10.1104/pp.105.073130 |
|
|
Ghazzawy H S, Bakr A, Mansour A T, et al. Paulownia trees as a sustainable solution for CO2 mitigation: assessing progress toward 2050 climate goals. Frontiers in Environmental Science, 2024, 12, 1307840.
doi: 10.3389/fenvs.2024.1307840 |
|
|
Ghosh R, Choi B, Cho B K, et al. Characterization of developmental- and stress-mediated expression of cinnamoyl-CoA reductase in kenaf (Hibiscus cannabinus L. ). The Scientific World Journal, 2014, 2014, 601845.
doi: 10.1155/2014/601845 |
|
|
Goicoechea M, Lacombe E, Legay S, et al. EgMYB2, a new transcriptional activator from Eucalyptus xylem, regulates secondary cell wall formation and lignin biosynthesis. The Plant Journal, 2005, 43 (4): 553- 567.
doi: 10.1111/j.1365-313X.2005.02480.x |
|
|
Goujon T, Ferret V, Mila I, et al. Down-regulation of the AtCCR1 gene in Arabidopsis thaliana: effects on phenotype, lignins and cell wall degradability. Planta, 2003, 217 (2): 218- 228.
doi: 10.1007/s00425-003-0987-6 |
|
|
Hou Y, Li X, Hu Y, et al. Drought weakens the positive effect of plant diversity on community biomass. Journal of Plant Ecology, 2024, 17 (4): rtae059.
doi: 10.1093/jpe/rtae059 |
|
|
Huang S, Kang X, Fu R, et al. Simultaneous down-regulation of dominant cinnamoyl CoA reductase and cinnamyl alcohol dehydrogenase dramatically altered lignin content in Mulberry. Plants, 2024, 13 (24): 3512.
doi: 10.3390/plants13243512 |
|
|
Jakubowski M. Cultivation potential and uses of Paulownia wood: A Review. Forests, 2022, 13 (5): 668.
doi: 10.3390/f13050668 |
|
|
Kawasaki T, Koita H, Nakatsubo T, et al. Cinnamoyl-CoA reductase, a key enzyme in lignin biosynthesis, is an effector of small GTPase Rac in defense signaling in rice. Proceedings of the National Academy of Sciences of the United States of America, 2006, 103 (1): 230- 235.
doi: 10.1073/pnas.0509875103 |
|
|
Krzywinski M, Schein J, Birol I, et al. Circos: an information aesthetic for comparative genomics. Genome Research, 2009, 19 (9): 1639- 1645.
doi: 10.1101/gr.092759.109 |
|
|
Lacombe E, Hawkins S, Van D J, et al. Cinnamoyl CoA reductase, the first committed enzyme of the lignin branch biosynthetic pathway: cloning, expression and phylogenetic relationships. The Plant Journal, 1997, 11 (3): 429- 441.
doi: 10.1046/j.1365-313x.1997.11030429.x |
|
|
Lauvergeat V, Lacomme C, Lacombe E, et al. Two cinnamoyl-CoA reductase (CCR) genes from Arabidopsis thaliana are differentially expressed during development and in response to infection with pathogenic bacteria. Phytochemistry, 2001, 57 (7): 1187- 1195.
doi: 10.1016/S0031-9422(01)00053-X |
|
| Leisner C P, Potnis N, Sanz-Saez A. Crosstalk and trade-offs: plant responses to climate change-associated abiotic and biotic stresses. Plant Cell and Environment, 2023, 46 (10): 2946- 2963. | |
|
Letunic I, Bork P. Interactive tree of life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Research, 2021, 49 (W1): W293- w296.
doi: 10.1093/nar/gkab301 |
|
|
Li J, Fan F, Wang L, et al. Cloning and expression analysis of cinnamoyl-CoA reductase (CCR) genes in sorghum. PeerJ, 2016, 4, e2005.
doi: 10.7717/peerj.2005 |
|
|
Ma Q H. Characterization of a cinnamoyl-CoA reductase that is associated with stem development in wheat. Journal of Experimental Botany, 2007, 58 (8): 2011- 2021.
doi: 10.1093/jxb/erm064 |
|
|
Ma Q H. Lignin biosynthesis and its diversified roles in disease resistance. Genes, 2024, 15 (3): 295.
doi: 10.3390/genes15030295 |
|
|
Mello B. Estimating timetrees with MEGA and the TimeTree resource. Molecular Biology and Evolution, 2018, 35 (9): 2334- 2342.
doi: 10.1093/molbev/msy133 |
|
|
Miedes E, Vanholme R, Boerjan W, et al. The role of the secondary cell wall in plant resistance to pathogens. Frontiers in Plant Science, 2014, 5, 358.
doi: 10.3389/fpls.2014.00358 |
|
|
Moura J C, Bonine C A, de Oliveira Fernandes Viana J, et al. Abiotic and biotic stresses and changes in the lignin content and composition in plants. Journal of Integrative Plant Biology, 2010, 52 (4): 360- 376.
doi: 10.1111/j.1744-7909.2010.00892.x |
|
|
Park S H, Mei C, Pauly M, et al. Downregulation of maize cinnamoyl-coenzyme a reductase via RNA interference technology causes brown midrib and improves ammonia fiber expansion-pretreated conversion into fermentable sugars for biofuels. Crop Science, 2012, 52 (6): 2687- 2701.
doi: 10.2135/cropsci2012.04.0253 |
|
|
Patzlaff A, McInnis S, Courtenay A, et al. Characterisation of a pine MYB that regulates lignification. The Plant Journal, 2003, 36 (6): 743- 754.
doi: 10.1046/j.1365-313X.2003.01916.x |
|
|
Piquemal J, Lapierre C, Myton K, et al. Down-regulation of cinnamoyl-CoA reductase induces significant changes of lignin profiles in transgenic tobacco plants. The Plant Journal, 1998, 13 (1): 71- 83.
doi: 10.1046/j.1365-313x.1998.00014.x |
|
|
Rejeb I B, Pastor V, Mauch-Mani B. Plant responses to simultaneous biotic and abiotic stress: molecular mechanisms. Plants, 2014, 3 (4): 458- 475.
doi: 10.3390/plants3040458 |
|
|
Sławińska N, Zając J, Olas B. Paulownia organs as interesting new sources of bioactive compounds. International Journal of Molecular Sciences, 2023, 24 (2): 1676.
doi: 10.3390/ijms24021676 |
|
|
Suzuki N, Rivero R M, Shulaev V, et al. Abiotic and biotic stress combinations. New Phytologist, 2014, 203 (1): 32- 43.
doi: 10.1111/nph.12797 |
|
|
Thévenin J, Pollet B, Letarnec B, et al. The simultaneous repression of CCR and CAD, two enzymes of the lignin biosynthetic pathway, results in sterility and dwarfism in Arabidopsis thaliana. Molecular Plant, 2011, 4 (1): 70- 82.
doi: 10.1093/mp/ssq045 |
|
|
Van A R, Leplé J C, Aerts D, et al. Improved saccharification and ethanol yield from field-grown transgenic poplar deficient in cinnamoyl-CoA reductase. Proceedings of the National Academy of Sciences of the United States of America, 2014, 111 (2): 845- 850.
doi: 10.1073/pnas.1321673111 |
|
|
Wang H, Tian W, Teng Y, et al. Rice cinnamoyl CoA reductase-like gene OsCCR14 involved in heat stress via regulation lignin and flavonoid accumulation. Plants, 2025, 14 (23): 3626.
doi: 10.3390/plants14233626 |
|
|
Xiong W, Reynolds M, Xu Y. Climate change challenges plant breeding. Current Opinion in Plant Biology, 2022, 70, 102308.
doi: 10.1016/j.pbi.2022.102308 |
|
|
Yin N, Li B, Liu X, et al. Two types of cinnamoyl-CoA reductase function divergently in accumulation of lignins, flavonoids and glucosinolates and enhance lodging resistance in Brassica napus. The Crop Journal, 2022, 10 (3): 647- 660.
doi: 10.1016/j.cj.2021.10.002 |
|
|
Zhan W, Cui L, Song N, et al. Comprehensive analysis of cinnamoyl-CoA reductase (CCR) gene family in wheat: implications for lignin biosynthesis and stress responses. BMC Plant Biology, 2025, 25 (1): 567.
doi: 10.1186/s12870-025-06605-8 |
|
|
Zhang H, Zhu J, Gong Z, et al. Abiotic stress responses in plants. Nature Reviews Genetics, 2022, 23 (2): 104- 119.
doi: 10.1038/s41576-021-00413-0 |
|
| Zhao Q, Dixon R A. 2011. Transcriptional networks for lignin biosynthesis: more complex than we thought? Trends in Plant Science, 16(4): 227−233. | |
|
Zhou R, Jackson L, Shadle G, et al. Distinct cinnamoyl CoA reductases involved in parallel routes to lignin in Medicago truncatula. Proceedings of the National Academy of Sciences of the United States of America, 2010, 107 (41): 17803- 17808.
doi: 10.1073/pnas.1012900107 |
|
|
Zoratti L, Karppinen K, Luengo E A, et al. Light-controlled flavonoid biosynthesis in fruits. Frontiers in Plant Science, 2014, 5, 534.
doi: 10.3389/fpls.2014.00534 |
| [1] | 赵佳玥,宗志洁,彭玺元,李志强,马星霞. 真菌降解木质素的酶及其代谢途径研究进展[J]. 林业科学, 2026, 62(3): 1-12. |
| [2] | 陈菲然,张子洁,方升佐. 纳米材料提升植物抗逆性的研究进展与林业应用展望[J]. 林业科学, 2025, 61(7): 83-93. |
| [3] | 姜思成,王妮,高会彬,周国强,杨海芸. 花叶矢竹矮化倒伏突变体DWF茎秆结构及细胞壁组分含量分析[J]. 林业科学, 2025, 61(3): 158-168. |
| [4] | 蒋康杰,吴文娟,黄丽菁,李家全,黎孔燕. 不同材种木质素影响底物吸附纤维素酶的水解机制[J]. 林业科学, 2024, 60(7): 140-148. |
| [5] | 熊怡心,王爽,马星霞,孙志勤. 木质素氧化酶系高产复合真菌培养体系构建[J]. 林业科学, 2024, 60(10): 133-142. |
| [6] | 邹春阳,吴文娟. 木质素结构单元对纤维素酶吸附的影响[J]. 林业科学, 2023, 59(6): 141-148. |
| [7] | 代琳心,王智辉,李振瑞,王佳军,刘杏娥,文甲龙,马建锋. 基于TG-FTIR的竹材细胞壁主要组分热解特性[J]. 林业科学, 2023, 59(11): 85-94. |
| [8] | 胥雅静,王佳伟,赵岩秋,江成,黄李超,安轶,曾为,张进,卢孟柱. 84K杨PagMSBP1/2a基因对木质素合成的影响[J]. 林业科学, 2022, 58(6): 56-65. |
| [9] | 桂有才,左宋林,金凯楠. 自发泡方法制备木质素基高比表面积泡沫炭[J]. 林业科学, 2022, 58(3): 139-148. |
| [10] | 李顺福,王慧敏,房丽莎,刘震. 短日照诱导白花泡桐顶芽死亡过程相关基因的表达[J]. 林业科学, 2022, 58(2): 148-158. |
| [11] | 贾茹,孙海燕,王玉荣,汪睿,赵荣军,任海青. 杉木无性系新品种‘洋020’和‘洋061’10年生幼龄材微观结构与力学性能的相关性[J]. 林业科学, 2021, 57(5): 165-175. |
| [12] | 贺学娇,楚立威,文爽爽,卢孟柱,唐芳. 以玉米为例探究单子叶植物重力响应及维管结构的变化[J]. 林业科学, 2021, 57(2): 93-102. |
| [13] | 杨昇,李改云. 金丝楸木材化学成分的不均一性[J]. 林业科学, 2021, 57(1): 169-177. |
| [14] | 吴文娟,邹春阳,黄丽菁,金永灿. 竹材在LiCl/DMSO溶剂体系中的溶解及再生性能[J]. 林业科学, 2020, 56(9): 201-206. |
| [15] | 周宇,韩雁明,李改云,储富祥. 酶解木质素质量浓度对纳米木质素粒子结构及载药行为的影响[J]. 林业科学, 2020, 56(3): 109-116. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||