

Scientia Silvae Sinicae ›› 2022, Vol. 58 ›› Issue (6): 66-78.doi: 10.11707/j.1001-7488.20220607
Previous Articles Next Articles
Xinyu Li1,Minqiu Wang1,Meiling Yuan1,Ueno Saneyoshi2,Xingtong Wu1,Mengying Cai1,3,Tsumura Yoshihiko3,Yafeng Wen1,*
Received:2021-07-17
Online:2022-06-25
Published:2022-09-24
Contact:
Yafeng Wen
CLC Number:
Xinyu Li,Minqiu Wang,Meiling Yuan,Ueno Saneyoshi,Xingtong Wu,Mengying Cai,Tsumura Yoshihiko,Yafeng Wen. Genetic Differentiation and Demographic History of Cryptomeria, A Relict Plant, in East Asia[J]. Scientia Silvae Sinicae, 2022, 58(6): 66-78.
Table 1
Locality information and sample size of Cryptomeria populations"
| 编号 NO. | 组群 Group | 种群 Population | 谱系 Lineage | 样本数量 Sample size | 样本来源 Location | 纬度 Latitude(N) | 经度 Longitude(E) | 海拔 Elevation/m |
| 1 | 柳杉 Cryptomeria japonica var. sinensis (CHN) | 黄山 Huangshan (WT) | CHS | 17 | 安徽黄山 Anhui Huangshan | 29.462 1° | 118.266 9° | 436 |
| 2 | 天目山 Tianmushan (XTM) | CHS | 23 | 浙江临安 Zhejiang Lin’an | 30.338 1° | 119.435 8° | 1 089 | |
| 3 | 天台山 Tiantaishan (TT) | CHS | 24 | 浙江天台 Zhejiang Tiantai | 29.2 495° | 121.0 948° | 897 | |
| 4 | 屏南 Pingnan (PNSL) | CHS | 23 | 福建屏南 Fujian Pingnan | 27.003 9° | 118.871 8° | 1 254 | |
| 5 | 福州 Fuzhou (FZ) | CHS | 21 | 福建福州 Fujian Fuzhou | 26.090 3° | 119.396 9° | 731 | |
| 6 | 天宝岩高海拔 Tianbaoyan high altitude (TB) | CHS | 10 | 福建永安 Fujian Yong’an | 25.918 3° | 117.557 5° | 1 500 | |
| 7 | 天宝岩 Tianbaoyan (TBY) | CHS | 20 | 福建永安 Fujian Yong’an | 25.965 6° | 117.512 3° | 1 132 | |
| 8 | 南平 Nanping (YTG) | CHS | 8 | 福建南平 Fujian Nanping | 26.738 4° | 118.146 8° | 846 | |
| 9 | 武夷山 Wuyishan (WYS) | CHS | 20 | 福建武夷山 Fujian Wuyishan | 27.759 8° | 117.693 6° | 892 | |
| 10 | 庐山 Lushan (LS) | LS | 23 | 江西庐山 Jiangxi Lushan | 29.549 7° | 115.966 8° | 895 | |
| 1 | 日本柳杉 Cryptomeria japonica var. japonica (JAN) | 青森 Ajigasawa (AJG) | Ura-sugi | 17 | 日本青森县 Japan Aomori | 40.675 6° | 140.205 3° | 319 |
| 2 | 秋田 Nibetsu (NBT) | Ura-sugi | 16 | 日本秋田县 Japan Akita | 39.806 1° | 140.260 0° | 315 | |
| 3 | 新泻 Donden (DND) | Ura-sugi | 16 | 日本新泻县 Japan Niigata | 38.139 7° | 138.383 3° | 725 | |
| 4 | 富山 Bijodaira (BJD) | Ura-sugi | 16 | 日本富山县 Japan Toyama | 36.576 1° | 137.458 9° | 628 | |
| 5 | 京都 Ashu (ASH) | Ura-sugi | 18 | 日本京都府 Japan Tokyo | 35.307 8° | 135.773 9° | 802 | |
| 6 | 宫崎 Oninome (ONN) | Ura-sugi | 6 | 日本宫崎县 Japan Miyazaki | 32.698 4° | 131.518 2° | 1 170 | |
| 7 | 鹿儿岛 Yakushima (YKU) | Omote-sugi | 7 | 日本鹿儿岛县 Japan Kagoshima | 30.303 5° | 130.573 1° | 1 071 | |
| 8 | 宫城 Ishinomaki (ISN) | Omote-sugi | 13 | 日本宫城县 Japan Miyagi-ken | 38.328 6° | 141.491 9° | 195 |
Table 2
Genetic diversity parameters of 14 nSSR loci"
| 序号 NO. | 位点 Locus | 等位基因数 Number of allele (Na) | 总的遗传变异 Total genetic diversity for the species (Ht) | 种群内遗传多样性 Genetic diversity within populations (Hs) | 种群间遗传分化系数 Genetic differentiation coefficient (FST) | 标准遗传分化系数 Gene differentiation factor (GST) | 多态性信息 含量Polymorphism information content (PIC) | 位点来源 Locus source |
| 1 | CS1895M | 29 | 0.715 | 0.664 | 0.093 | 0.072 | 0.594 | |
| 2 | BY900902 | 7 | 0.636 | 0.592 | 0.063 | 0.069 | 0.516 | |
| 3 | Cjgssr31 | 33 | 0.919 | 0.857 | 0.071 | 0.067 | 0.798 | |
| 4 | BY894091 | 3 | 0.038 | 0.038 | 0.010 | 0.003 | 0.034 | |
| 5 | CS1737M | 2 | 0.499 | 0.449 | 0.096 | 0.100 | 0.337 | |
| 6 | Cjgssr175 | 20 | 0.644 | 0.493 | 0.231 | 0.235 | 0.417 | Moriguchi Y. et al., (2005) |
| 7 | Cjgssr7 | 22 | 0.774 | 0.672 | 0.150 | 0.132 | 0.588 | |
| 8 | BY896143 | 3 | 0.315 | 0.290 | 0.068 | 0.079 | 0.234 | |
| 9 | BY898881 | 11 | 0.693 | 0.624 | 0.095 | 0.099 | 0.547 | |
| 10 | Cjgssr120 | 26 | 0.604 | 0.482 | 0.201 | 0.202 | 0.441 | |
| 11 | Cjgssr181 | 31 | 0.635 | 0.560 | 0.103 | 0.118 | 0.481 | |
| 12 | BJ939490 | 5 | 0.313 | 0.281 | 0.123 | 0.102 | 0.236 | |
| 13 | CS1522M | 18 | 0.597 | 0.541 | 0.098 | 0.094 | 0.490 | |
| 14 | BY909057 | 9 | 0.597 | 0.562 | 0.060 | 0.058 | 0.458 | |
| 均值 | Mean | 15.643 | 0.570 | 0.508 | 0.112 | 0.109 | 0.441 |
Table 3
Genetic diversity parameters of the Cryptomeria populations"
| 种群 Population | 等位基因数 Number of alleles(Na) | 观测杂合度 Observed heterozygosity (Ho) | 期望杂合度 Expected heterozygosity (He) | 近交系数 Inbreeding coefficient (FIS) | 等位基因丰富度 Allelic richness (Ar) | 私有等位基因丰富度 Private alleles richness (PAr) |
| 黄山Huangshan (WT) | 3.357 | 0.303 | 0.412 | 0.294 | 2.636 | 0.148 |
| 天目山Tianmushan (XTM) | 4.000 | 0.508 | 0.491 | -0.012 | 2.826 | 0.125 |
| 天台山Tiantaishan (TT) | 4.643 | 0.464 | 0.511 | 0.113 | 3.083 | 0.209 |
| 屏南Pingnan (PNSL) | 4.143 | 0.493 | 0.502 | 0.041 | 3.111 | 0.125 |
| 福州Fuzhou (FZ) | 4.643 | 0.439 | 0.486 | 0.121 | 3.158 | 0.143 |
| 天宝岩高海拔Tianbaoyan high altitude (TB) | 2.714 | 0.293 | 0.360 | 0.236 | 2.453 | 0.144 |
| 天宝岩Tianbaoyan (TBY) | 5.143 | 0.461 | 0.512 | 0.125 | 3.360 | 0.233 |
| 南平Nanping (YTG) | 2.857 | 0.439 | 0.431 | 0.050 | 2.752 | 0.065 |
| 武夷山Wuyishan (WYS) | 4.929 | 0.446 | 0.499 | 0.131 | 3.409 | 0.202 |
| 庐山Lushan (LS) | 9.286 | 0.575 | 0.695 | 0.194 | 5.271 | 0.976 |
| 中国平均值CHN Mean | 4.571 | 0.442 | 0.490 | 0.123 | 3.206 | 0.240 |
| 青森Ajigasawa (AJG) | 4.500 | 0.471 | 0.454 | -0.006 | 3.216 | 0.089 |
| 秋田Nibetsu (NBT) | 4.643 | 0.477 | 0.493 | 0.064 | 3.364 | 0.072 |
| 新泻Donden (DND) | 4.929 | 0.463 | 0.448 | -0.002 | 3.239 | 0.200 |
| 富山Bijodaira (BJD) | 4.786 | 0.536 | 0.493 | -0.055 | 3.316 | 0.217 |
| 京都Ashu (ASH) | 5.714 | 0.477 | 0.492 | 0.060 | 3.535 | 0.293 |
| 宫崎Oninome (ONN) | 3.071 | 0.536 | 0.450 | -0.100 | 3.071 | 0.107 |
| 鹿儿岛Yakushima (YKU) | 4.357 | 0.611 | 0.528 | -0.080 | 4.073 | 0.684 |
| 宫城Ishinomaki (ISN) | 5.071 | 0.511 | 0.529 | 0.073 | 3.672 | 0.284 |
| 日本平均值JAN Mean | 4.634 | 0.510 | 0.486 | 0.009 | 3.436 | 0.243 |
| 总平均值Total Mean | 4.599 | 0.472 | 0.488 | 0.060 | 3.308 | 0.240 |
Table 4
Analysis of molecular variance (AMOVA) of Cryptomeria populations"
| 变异来源 Source of variation | 自由度 df | 方差总和 Sum of squares | 变异组分 Variance components | 变异百分比 Percentage of variation (%) | P |
| 组间 Among groups | 1 | 70.972 | 0.199 | 4.85 | < 0.001 |
| 种群间 Among populations | 16 | 246.363 | 0.361 | 8.78 | < 0.001 |
| 种群内 Within populations | 578 | 2 052.437 | 3.551 | 86.37 | < 0.001 |
| 总计 Total | 595 | 2 369.772 | 4.092 | 100.00 | - |
| FST=0.136 Nm=1.588 | |||||
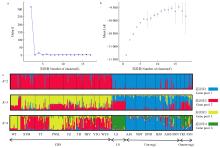
Fig.2
Results of STRUCTURE analysis of the Cryptomeria populations a: Relationship between Delta K and K value; b: Relationship between Mean LnK and K value; c: Histogram of the STRUCTURE analysis for the model with K=2, K=3 and K=4;; CHS: the populations of Southeastern China; LS: the populations of Lushan China; Ura-sugi: the populations of the Japan Sea coast; Omote-sugi: the populations of the Pacific coast."
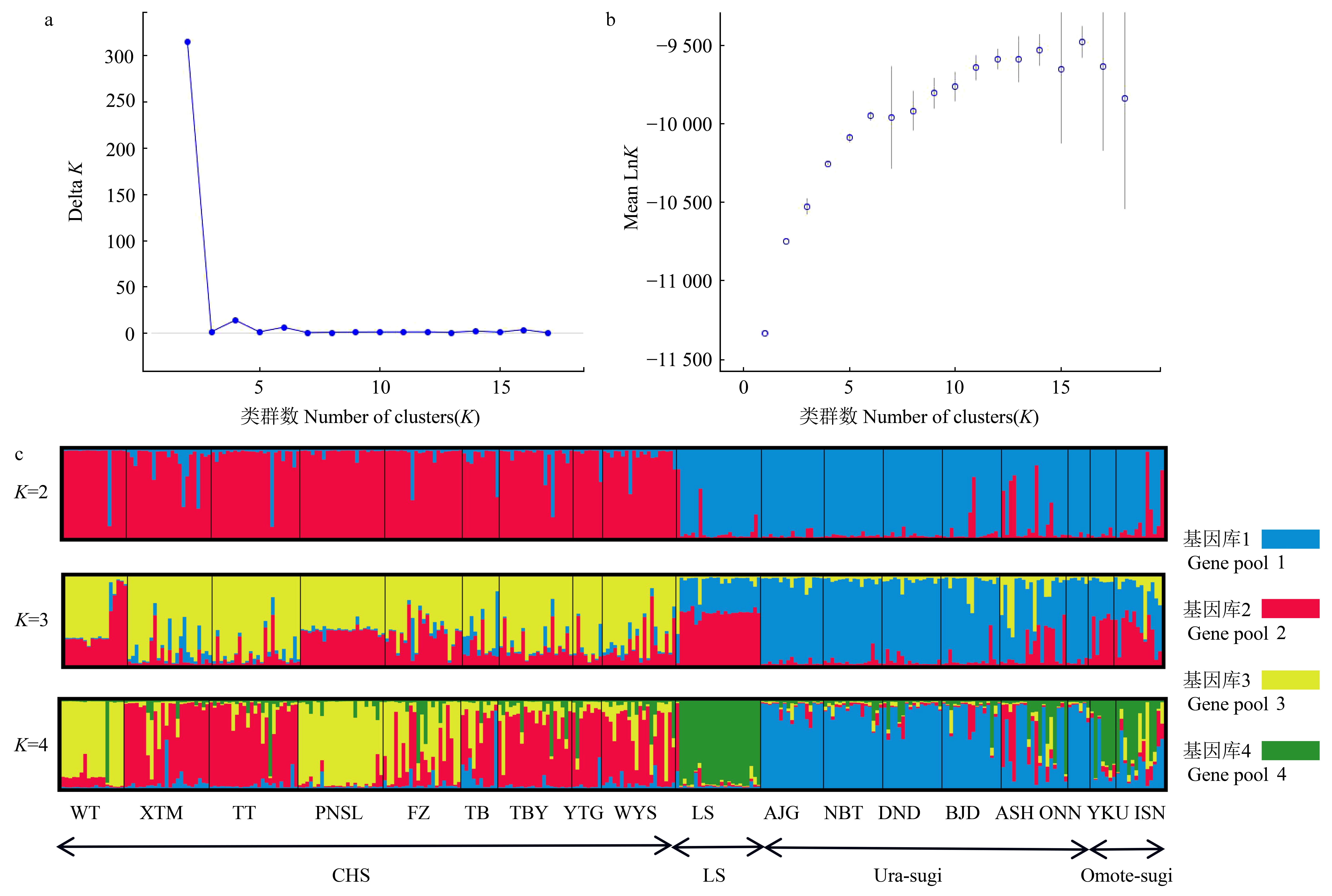

Fig.7
The scenarios of Cryptomeria populations based on Approximate Bayesian Computation (K = 4) CHS: the populations of Southeastern China; LS: the populations of Lushan China; Ura-sugi: the populations of the Japan Sea coast; Omote-sugi: the populations of the Pacific coast; t1, t2, t3: the generation of scenario; N1, N2, N3, N4, NA: the size of effective population."
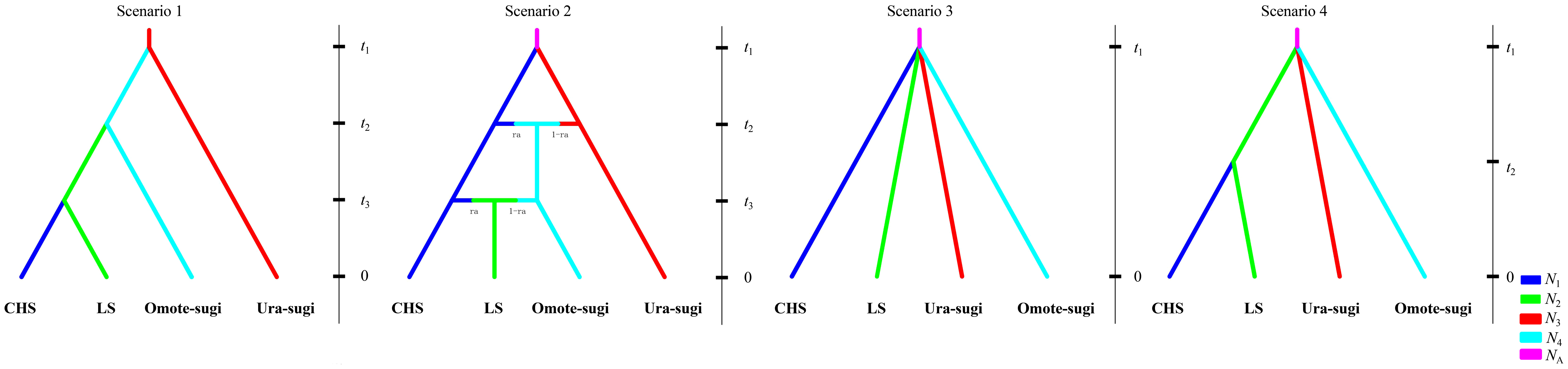
Table 5
Median estimation of posterior distributions for each scenario based on DIY ABC (K = 4)"
| 事件1 Scenario 1 | 事件2 Scenario 2 | 事件3 Scenario 3 | 事件4 Scenario 4 | |
| 后验概率Posterior probability (PP) | 0.000 8 | 0.000 0 | 0.991 9 | 0.007 3 |
| 种群大小Effective population size (N1) | 4 490 | 6 930 | 3 820 | 4 690 |
| 种群大小Effective population size (N2) | 8 870 | 9 700 | 7 840 | 8 800 |
| 种群大小Effective population size (N3) | 2 600 | 5 550 | 4 850 | 4 740 |
| 种群大小Effective population size (N4) | 2 950 | 6 870 | 7 350 | 7 190 |
| 种群大小Effective population size (NA) | - | 22.5 | 44.6 | 42.3 |
| 代数Generation (t1) | 720 | 2 540 | 1 140 | 1 450 |
| 代数Generation (t2) | 559 | 1 840 | - | 1 360 |
| 代数Generation (t3) | 205 | 1 790 | - | - |
| 程小毛, 林开文, 姜永雷, 等. 玉龙雪山不同海拔急尖长苞冷杉遗传多样性分析. 江西农业大学学报, 2016, 38 (4): 654- 659. | |
| Cheng X M , Lin K W , Jiang Y L , et al. Genetic diversity of Abies georgei var.smithii at different altitudes in Jade Dragon Snow Mountain. Acta Agriculturae Universitatis Jiangxiensis, 2016, 38 (4): 654- 659. | |
| 李江伟. 2014. 基于NSSR和CPSSR标记的台湾杉遗传多样性研究. 武汉: 华中农业大学. | |
| Li J W. 2014. Genetic diversity of Taiwania cryptomerioides based on nSSR and cpSSR molecular markers. Wuhan: Huazhong Agricultural University. [in Chinese] | |
|
李霞, 王利宝, 文亚峰, 等. 杉木不同世代育种群体的遗传多样性. 林业科学, 2020, 56 (11): 53- 61.
doi: 10.11707/j.1001-7488.20201106 |
|
|
Li X , Wang L B , Wen Y F , et al. Genetic diversity of Chinese Fir (Cunninghamia lanceolata) breeding populations among different generations. Scientia Silvae Sinicae, 2020, 56 (11): 53- 61.
doi: 10.11707/j.1001-7488.20201106 |
|
| 刘建全. "整合物种概念"和"分化路上的物种". 生物多样性, 2016, 24 (9): 1004- 1008. | |
| Liu J Q . "The integrative species concept" and "species on the speciation way". Biodiversity Science, 2016, 24 (9): 1004- 1008. | |
| 邱英雄, 鹿启祥, 张永华, 等. 东亚第三纪孑遗植物的亲缘地理学: 现状与趋势. 生物多样性, 2017, 25 (2): 136- 146. | |
| Qiu Y X , Lu Q X , Zhang Y H , et al. Phylogeography of East Asia's tertiary relict plants: current progress and future prospects. Biodiversity Science, 2017, 25 (2): 136- 146. | |
|
沈作奎, 鲁胜平. 日本柳杉合理经营密度的研究. 湖北民族学院学报(自然科学版), 2004, 22 (4): 57- 59.
doi: 10.3969/j.issn.1008-8423.2004.04.018 |
|
|
Shen Z K , Lu S P . Study on the rational density for Cryptomoria japonica plantation. Journal of Hubei Institute for Nationalities (Natural Science Edition), 2004, 22 (4): 57- 59.
doi: 10.3969/j.issn.1008-8423.2004.04.018 |
|
|
王江, 刘军, 黄永强, 等. 柳杉起源及天然分布. 四川林业科技, 2007, 28 (4): 92- 94.
doi: 10.3969/j.issn.1003-5508.2007.04.021 |
|
|
Wang J , Liu J , Huang Y Q , et al. The origin and natural distribution of Cryptomeria. Journal of Sichuan Forestry Science and Technology, 2007, 28 (4): 92- 94.
doi: 10.3969/j.issn.1003-5508.2007.04.021 |
|
|
文亚峰, 韩文军, 吴顺. 植物遗传多样性及其影响因素. 中南林业科技大学学报: 自然科学版, 2010, 30 (12): 80- 87.
doi: 10.3969/j.issn.1673-923X.2010.12.016 |
|
|
Wen Y F , Han W J , Wu S . Plant genetic diversity and its influencing factors. Journal of Central South University of Forestry Science & Technology: Natural Science Edition, 2010, 30 (12): 80- 87.
doi: 10.3969/j.issn.1673-923X.2010.12.016 |
|
| 于永福. 杉科植物的起源、演化及其分布. 植物分类学报, 1995, 33 (4): 362- 389. | |
| Yu Y F . Origin evolution and distribution of the Taxodiacea. Journal of Systematics and Evolution, 1995, 33 (4): 362- 389. | |
| 袁美灵, 文亚峰, 武星彤, 等. 柳杉遗传资源及其研究进展. 四川林业科技, 2019, 40 (5): 91- 95. | |
| Yuan M L , Wen Y F , Wu X T , et al. Genetic resources and research progress of Cryptomeria. Journal of Sichuan Forestry Science and Technology, 2019, 40 (5): 91- 95. | |
| 张景丽. 2014. 柳杉优树资源的遗传多样性分析及杂交亲本的筛选. 杭州: 浙江农林大学. | |
| Zhang J L. 2014. The analysis on genetic diversity of superior Cryptomehia fortunei resources and screening of hybrid parent. Hangzhou: Zhejiang A & F University. [in Chinese] | |
| Begon M, Townsend C R, Harper J L. 2006. Ecology: from individuals to ecosystems. Oxford: Blackwell. | |
| Bennett K . Evolution and ecology: the pace of life. Cambridge: Cambridge University Press, 1997. | |
|
Cai M Y , Wen Y F , Uchiyama K , et al. Population genetic diversity and structure of ancient tree populations of Cryptomeria japonica var.sinensis based on RAD-seq data. Forests, 2020, 11, 1192.
doi: 10.3390/f11111192 |
|
|
Cao Y N , Comes H P , Sakaguchi S , et al. Evolution of East Asia's arcto-tertiary relict Euptelea (Eupteleaceae) shaped by late neogene vicariance and quaternary climate change. BMC Evolutionary Biology, 2016, 16 (1): 66.
doi: 10.1186/s12862-016-0636-x |
|
|
Chen Y , Yang S Z , Zhao M S , et al. Demographic genetic structure of Cryptomeria japonica var.sinensis in Tianmushan nature reserve, China. Journal of Integrative Plant Biology, 2008, 50 (9): 1171- 1177.
doi: 10.1111/j.1744-7909.2008.00725.x |
|
|
Chen Y S , Deng T , Zhou Z , et al. Is the East Asian flora ancient or not?. National Science Review, 2018, 5, 920- 932.
doi: 10.1093/nsr/nwx156 |
|
|
Cornuet J M , Pierre P , Julien V , et al. DIYABC v2.0: a software to make approximate Bayesian computation inferences about population history using single nucleotide polymorphism, DNA sequence and microsatellite data. Bioinformatics, 2014, 30 (8): 1187- 1189.
doi: 10.1093/bioinformatics/btt763 |
|
|
Earl D A , Von Holdt B M . STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conservation Genetics Resources, 2012, 4 (2): 359- 361.
doi: 10.1007/s12686-011-9548-7 |
|
| Excoffier L , Laval G , Schneider S . Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evolutionary Bioinformatics Online, 2005, 1 (4A): 47- 50. | |
| Fu L G, Li N, Mill R. 1999. Cryptomeria. In: Flora of China. Beijing: Science Press, 56-57. | |
|
Goudet J . FSTAT (version 1.2) : a computer program to calculate F-statistics. Journal of Heredity, 1995, 86 (6): 485- 486.
doi: 10.1093/oxfordjournals.jhered.a111627 |
|
|
Harrison S P , Yu G , Takahara H , et al. Diversity of temperate plants in East Asia. Nature, 2001, 413 (6852): 129- 130.
doi: 10.1038/35093166 |
|
| Hayashi Y. 1960. Taxonomical and phytogeographical study of Japanese conifers. Tokyo, Japan: Norin-Shuppan. | |
|
Hewitt G M . Some genetic consequences of ice ages, and their role, in divergence and speciation. Biological Journal of the Linnean Society, 1996, 58 (3): 247- 276.
doi: 10.1006/bijl.1996.0035 |
|
|
Hewitt G M . Genetic consequences of climatic oscillations in the quaternary. Philosophical Transactions of the Royal Society of London, 2004, 359 (1442): 183- 195.
doi: 10.1098/rstb.2003.1388 |
|
|
Hohmann N , Wolf E M , Rigault P , et al. Ginkgo biloba's footprint of dynamic Pleistocene history dates back only 390, 000 years ago. BMC Genomics, 2018, 19 (1): 299.
doi: 10.1186/s12864-018-4673-2 |
|
| Hulce D , Li X , SnyderLeiby T , et al. GeneMarker genotyping software: tools to increase the statistical power of DNA fragment analysis. Journal of Biomolecular Techniques, 2011, 22 (Suppl): S35- S36. | |
|
Jakobsson M , Rosenberg N A . CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics, 2007, 23 (14): 1801- 1806.
doi: 10.1093/bioinformatics/btm233 |
|
|
Jombart T . adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics (Oxford, England), 2008, 24 (11): 1403- 1405.
doi: 10.1093/bioinformatics/btn129 |
|
|
Kalinowski S . HP-RARE 1.0: A computer program for performing rarefaction on measures of allelic richness. Molecular Ecology Notes, 2005, 5 (1): 187- 189.
doi: 10.1111/j.1471-8286.2004.00845.x |
|
| Kimura M K , Kabeya D , Saito T , et al. Effects of genetic and environmental factors on clonal reproduction in old-growth natural populations of Cryptomeria japonica. Forest Ecology & Management, 2013, 304 (Complete): 10- 19. | |
|
Kimura M K , Uchiyama K , Nakao K , et al. Evidence for cryptic northern refugia in the last glacial period in Cryptomeria japonica. Annals of Botany, 2014, 114 (8): 1687- 1700.
doi: 10.1093/aob/mcu197 |
|
|
Kumar S , Stecher G , Tamura K . MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molecular Biology and Evolution, 2016, 33 (7): 1870- 1874.
doi: 10.1093/molbev/msw054 |
|
| Kojo Y . A dendrochronological study of Cryptomeria japonica in Japan. Tree Ring Bulletin, 1987, 47, 1- 21. | |
| Ledig F T. 1998. Genetic variation in Pinus. In: Ecology and Biogeography of Pinus. Cambridge: Cambridge University Press, 251-280. | |
| Milne R I , Abbott R J . The origin and evolution of tertiary relict flora. Advances in Botanical Research, 2002, 38, 281- 314. | |
|
Moriguchi Y , Iwata H , Ujino-Ihara T , et al. Development and characterization of microsatellite markers for Cryptomeria japonica D.Don. Theoretical and Applied Genetics, 2003, 106 (4): 751- 758.
doi: 10.1007/s00122-002-1149-0 |
|
|
Moriguchi Y , Ueno S , Higuchi Y , et al. Establishment of a microsatellite panel covering the sugi (Cryptomeria japonica) genome, and its application for localization of a male-sterile gene (ms-2). Molecular Breeding, 2014, 33 (2): 315- 325.
doi: 10.1007/s11032-013-9951-8 |
|
|
Moriguchi N , Uchiyama K , Miyagi R , et al. Inferring the demographic history of Japanese cedar, Cryptomeria japonica, using amplicon sequencing. Heredity, 2019, 123 (3): 371- 383.
doi: 10.1038/s41437-019-0198-y |
|
|
Peakall R , Smouse P E . GenAlEx version 6.5: Genetic analysis in Excel. Population genetic software for teaching and research-An update. Bioinformatics, 2012, 28 (19): 2537- 2539.
doi: 10.1093/bioinformatics/bts460 |
|
|
Pritchard J K , Stephens M , Donnelly P . Inference of population structure using multilocus genotype data. Genetics, 2000, 155 (2): 945- 959.
doi: 10.1093/genetics/155.2.945 |
|
| Qi X S , Yuan N , Comes H P , et al. A strong 'filter' effect of the East China sea land bridge for East Asia's temperate plant species: inferences from molecular phylogeography and ecological niche modelling of Platycrater arguta (Hydrangeaceae). BMC Evolutionary Biology, 2014, 14 (1): 41. | |
| Qiu Y X , Sun Y , Zhang X P , et al. Molecular phylogeography of East Asian Kirengeshoma (Hydrangeaceae) in relation to Quaternary climate change and landbridge configurations. New Phytologist, 2009, 183 (2): 480- 495. | |
| Rosenberg N A . DISTRUCT: A program for the graphical display of population structure. Molecular Ecology Notes, 2004, 4 (1): 137- 138. | |
| Tani N , Takahashi T , Iwata H , et al. A consensus linkage map for sugi (Cryptomeria japonica) from two pedigrees, based on microsatellites and expressed sequence tags. Genetics, 2003, 165 (3): 1551- 1568. | |
| Tiffney B H . The Eocene North Atlantic land bridge: its importance in tertiary and modern phytogeography of the northern hemisphere. Journal of the Arnold Arboretum, 1985, 66 (2): 243- 273. | |
| Tiffney B H , Manchester S . The use of geological and paleontological evidence in evaluating plant phylogeographic hypotheses in the northern hemisphere tertiary. International Journal of Plant Sciences, 2001, 162 (6 Suppl): S3- S17. | |
| Tsumura Y , Kado T , Takahashi T , et al. Genome scan to detect genetic structure and adaptive genes of natural populations of Cryptomeria japonica. Genetics, 2007, 176 (4): 2393- 2403. | |
| Tsumura Y. 2011. Cryptomeria. In Wild Crop Relatives: Genomic and Breeding Resources: Forest Trees. Germany, Berlin: Springer, 49-63. | |
| Tsumura Y , Uchiyama K , Moriguchi Y , et al. Genome scanning for detecting adaptive genes along environmental gradients in the Japanese conifer, Cryptomeria japonica. Heredity, 2012, 109 (6): 349- 360. | |
| Tsumura Y , Uchiyama K , Moriguchi Y , et al. Genetic differentiation and evolutionary adaptation in Cryptomeria japonica. G3: Genes, Genomes, Genetics, 2014, 4 (12): 2389- 2402. | |
| Tsumura Y , Kimura M , Nakao K , et al. Effects of the last glacial period on genetic diversity and genetic differentiation in Cryptomeria japonica in East Asia. Tree Genetics & Genomes, 2020, 16 (1): 19. | |
| Wang I J , Glor R E , and Losos J B . Quantifying the roles of ecology and geography in spatial genetic divergence. Ecology Letters, 2013, 16 (2): 175- 182. | |
| Wu X T , Ruhsam M , Wen Y F , et al. The last primary forests of the tertiary relict Glyptostrobus pensilis contain the highest genetic diversity. Forestry, 2019, 93 (3): 359- 375. | |
| Yuan N , Sun Y , Comes H P , et al. Understanding population structure and historical demography in a conservation context: population genetics of the endangered Kirengeshoma palmata (Hydrangeaceae). American Journal of Botany, 2014, 101 (3): 521- 529. |
| [1] | Wenting Pan,Jianjun Sun,Qinqin Yuan,Lili Zhang,Kangqiao Deng,Yueqiao Li. Analysis of Genetic Diversity and Structure in Different Provenances of Liriodendron by RAD-seq Technique [J]. Scientia Silvae Sinicae, 2022, 58(4): 74-81. |
| [2] | Yuanchen Zhang,Shaohui Lu,Dongfeng Gong,Xingli Ma. Analysis of Genetic Diversity and Genetic Structure in Geographic Populations of Corythucha ciliata(Hemiptera: Tingidae) from China Based on Mitochondrial DNA COⅠ Gene Sequences [J]. Scientia Silvae Sinicae, 2021, 57(8): 102-111. |
| [3] | Huanhuan Chen,Gexi Xu,Fanqiang Ma,Shun Liu,Miaomiao Zhang,Xiangwen Cao,Jian Chen,Guangdong Zhao,Hongguo Yang,Zuomin Shi. Phylogenetic Structure of Undergrowth Layers across Subalpine Dark Coniferous Forests and Their Post-Harvesting Secondary Forests in Western Sichuan [J]. Scientia Silvae Sinicae, 2020, 56(7): 1-11. |
| [4] | Yihong Meng,Gangbiao Xu,Mengzhu Lu,Xiaolong Jiang,Feilong Guo. Population Genetic Structure and Demographic History of Disanthus cercidifolius var. longipes [J]. Scientia Silvae Sinicae, 2020, 56(7): 55-62. |
| [5] | Xia Li,Libao Wang,Yafeng Wen,Jun Lin,Xingtong Wu,Meiling Yuan,Yuan Zhang,Minqiu Wang,Xinyu Li. Genetic Diversity of Chinese Fir (Cunninghamia lanceolata) Breeding Populations among Different Generations [J]. Scientia Silvae Sinicae, 2020, 56(11): 53-61. |
| [6] | Ma Songmei, Wang Chuncheng, Sun Fangfang, Wei Bo, Nie Yingbin. Genetic Diversity of an Endangered Plant Amygdalus ledebouriana in Xinjiang [J]. Scientia Silvae Sinicae, 2019, 55(9): 71-80. |
| [7] | Zhang Shouke, Fang Linxin, Liu Yaning, Wang Yi, Zhang Wei, Shu Jinping, Zhang Yabo, Wang Yangdong, Wang Haojie. Genetic Differentiation and Structural Variation of ATP Synthase Gene of Curculio chinensis (Coleptera: Curculionidae) under Selection Pressure at Different Altitudes [J]. Scientia Silvae Sinicae, 2019, 55(6): 65-73. |
| [8] | Hao Minhui, Li Xiaoyu, Xia Mengjie, He Huaijiang, Zhang Chunyu, Zhao Xiuhai. Effects of Tending Felling on Functional and Phylogenetic Structures in a Multi-Species Temperate Secondary Forest at Jiaohe in Jilin Province [J]. Scientia Silvae Sinicae, 2018, 54(5): 1-9. |
| [9] | Bao Wenquan, Wuyun Tana, Du Hongyan, Li Tiezhu, Liu Huimin, Wang Lin, Bai Yu. Genetic Diversity and Population Structure of Amygdalus mira in the Tibet Plateau in China Based on SSR Markers [J]. Scientia Silvae Sinicae, 2018, 54(2): 30-41. |
| [10] | Yang Hanbo, Zhang Rui, Wang Bangshun, Xu Zhaoyou, Chen Huanwei, Zhou Zhichun. Analysis of Genetic Diversity in Schima superba Plus Tree Germplasms by SSR Markers [J]. Scientia Silvae Sinicae, 2017, 53(5): 43-53. |
| [11] | Xu Jin, Zhu Chenchen, Ouyang Lei, Shi Jisen. The Difference in Regeneration Capacity of Selected Cryptomeria Group Clones in vitro [J]. Scientia Silvae Sinicae, 2016, 52(8): 46-52. |
| [12] | Li He, Li Yang, Xu Jianping, Zhou Guoying. Population Genetic Structure of Colletotrichum fructicola from Oil-Tea and Other Host Plants in Hainan province [J]. Scientia Silvae Sinicae, 2016, 52(10): 80-88. |
| [13] | Li Pei, Que Qingmin, Ouyang Kunxi, Li Juncheng, Zhan Xin, Zhu Qin, Zhang Junjie, Deng Xiaomei, Chen Xiaoyang. Genetic Diversity of Toona ciliata from Different Provenances Based on Sequence-Related Amplified Polymorphism (SRAP) Markers [J]. Scientia Silvae Sinicae, 2016, 52(1): 62-70. |
| [14] | Wang Yanhong, Yu Qi, Yang Jia, Zhao Peng, Li Zhonghu, Zhao Guifang. Genetic Diversity and Population Structure of Quercus serrata var.brevipetiolata Revealed by nSSR Markers [J]. Scientia Silvae Sinicae, 2015, 51(12): 121-131. |
| [15] | Wen Yafeng, Han Wenjun, Zhou Hong, Xu Gangbiao. SSR Mining and Development of EST-SSR Markers for Cunninghamia lanceolata Based on Transcriptome Sequences [J]. Scientia Silvae Sinicae, 2015, 51(11): 40-49. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||