

Scientia Silvae Sinicae ›› 2020, Vol. 56 ›› Issue (11): 62-72.doi: 10.11707/j.1001-7488.20201107
Previous Articles Next Articles
Yuan Li1,Jinhuan Chen2,Zhao Jin1,Jingya Hou1,Yusong Jiang1,Haitao Xing1,*
Received:2020-06-22
Online:2020-11-25
Published:2020-12-30
Contact:
Haitao Xing
CLC Number:
Yuan Li,Jinhuan Chen,Zhao Jin,Jingya Hou,Yusong Jiang,Haitao Xing. Functions of NAC128 Gene from Populus trichocarpa in Secondary Cell Wall Formation[J]. Scientia Silvae Sinicae, 2020, 56(11): 62-72.
Table 1
List of primers"
| 基因名称 Gene name | 正向引物序列 Forward primer sequence (5′—3′) | 反向引物序列 Reverse primer sequence (5′—3′) |
| 克隆和载体构建引物Primers for cloning and vector construction | ||
| PtrNAC128 | ATGACTACTAAGAGTAATATGGCTTCC | GACCAACCCATGATGATCCTGGTTGTC |
| PtrNAC128 (BglⅡ, SpeⅠ) | GCGAGATCTGATGACTACTAAGAGTAATATGGCTTCC | GGACTAGTGACCAACCCATGATGATCCTGGTTGTC |
| 定量RT-PCR引物Primers for qRT-PCR | ||
| PtrNAC128 | GGATAAGAGAGGCTTCAGTAGC | TCAAGAATGATGGAGGTGTG |
| PtoCCOAOMT1 | CAAGAGGTTGATTGAGCTTG | GGTCAGCAGCAAGTGCCTTG |
| PtoCCR2 | CTGTTCAAGCTTATGTGCATG | GTGGAGAACGCTCTCAGAGC |
| PtoCOMT2 | CATGAAGTGGATATGCCATG | GTTGAATGCACAGCACATTAC |
| PtoC3H3 | GAGGTTCCTGGAGGAGGATG | GGAGTCGTCATGTAAGTGAC |
| PtoPAL4 | CCTACATTGACGATCCTTGCAG | GACCTGCATTCCTTGATCCTG |
| PtoHCT1 | ATCAGCATGTAAGGCACGCGG | TGCCAAAGTAACCAGGTGGAAGCGT |
| PtoCAD1 | CAAGCTGATCTTGATGGGTG | CGAATCTATATCTCACATC |
| Ptr4CL5 | CATCCGAGGTGATCAGATCATG | CACAGCAGCATCAGATATCC |
| PtrF5H2 | GAGTCCAGCAAGAGCTCGCAG | GCATAAGCATTGATCATCAC |
| PtoCesA2B | AGGTTAAGATGGAGCGG | ACGAGGTTGATGATCAAGCC |
| PtoCesA3A | CGGATATGGATCATGGGGTCC | GGATAGAGATGGACAATGAC |
| PtoGT8D | GTGCCTGGGCTTATGGCATG | CTAGCCAAGGCTTTGCTCGACC |
| PtoGT43B | CTCAATCCTCTGGGATCCTG | GTCTCGTCTTCAAGAGCTAC |
| PtoGT43D | GGTAGAGCCACTTGGGAGC | CACAAGGCAGTATCTCTG |
| PtoCSE1 | GACAGTCCACGGCACGGCTG | CCTCTCAACCCTCTCGTC |
| PtoMYB192 | CGTGACCGAAATCCCATTCGAG | GAGTGGCATGTCATGCTG |
| PtoMYB161 | CGAGCAGAAACCTTCCTTCTC | CCAGCCCTGCTCTTAGGC |
| PtoNAC156 | CCACTCTTGTCGAGTAC | CTTCTCAATGTATCCTGCC |
| PtoMYB152 | GAAGACTTGCTACTGCCAGAT | TCATTCTTGAGCACTGATTG |
| PtoMYB028 | CGTGGTAACTTCAGCAAAGAG | GAGGAGGTTGAGGAGGATTC |
| PtoMYB090 | GAGTTGGTTGAGAAGTATGGC | CGGTGAGAAGCAAGAAGTCT |
| PtoMYB175 | GCTGTGAAGAACCATTGGC | TGCTCATTGACTAACAACGC |
| PtoNAC105 | GGATAACTTGCCCTTCTTGTG | CACCTTTGCTTCTAAATGCC |
| PtoNAC154 | CAAGACCAGGCAAGAATCC | GATGGAAGAAGTGGCGAA |
| Pto18S | GGCATGGAAGGTGATGCAGATC | CTGTGTCAAACAAGAACTTGTCC |
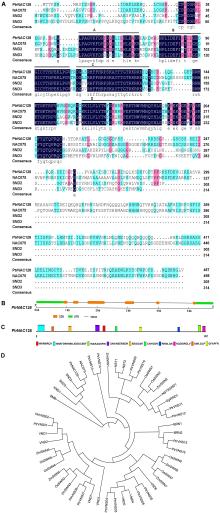
Fig.1
Comparison of PtrNAC128 with other NAC amino acid sequences A. Multiple sequence alignment between PtrNAC128 and the 3 homologes of Arabidopsis (Identical amino acids are shaded in dark blue, the conserved NAC domain are overlined); B. Gene structure of PtrNAC128; C. Structure of PtrNAC128 protein potential motifs; D. Phylogenetic analysis of PtrNAC128 and other NAC proteins by the neighbor-joining method using MEGA version X. Ptr: Populus trichocarpa; Os: Oryza sativa; Zm: Zea mays; Eg: Eucalyptus grandis; Others: Arabidopsis thaliana."

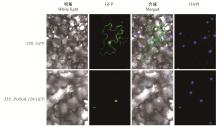
Fig.3
Subcellular localization analysis of PtrNAC128 Confocal images of localization of PtrNAC128-GFP in tobacco(Nicotiana benthamiana) leaf epidermal cells. Bright field and fluorescent micrographs show the nuclear localization of PtrNAC128-GFP and free GFP expressed both in nuclear and cytomembrane localization in tobacco leaf epidermal cells. DAPI (40, 6-diamidino-2-phenylindole dihydrochloride), a nuclear staining dye."
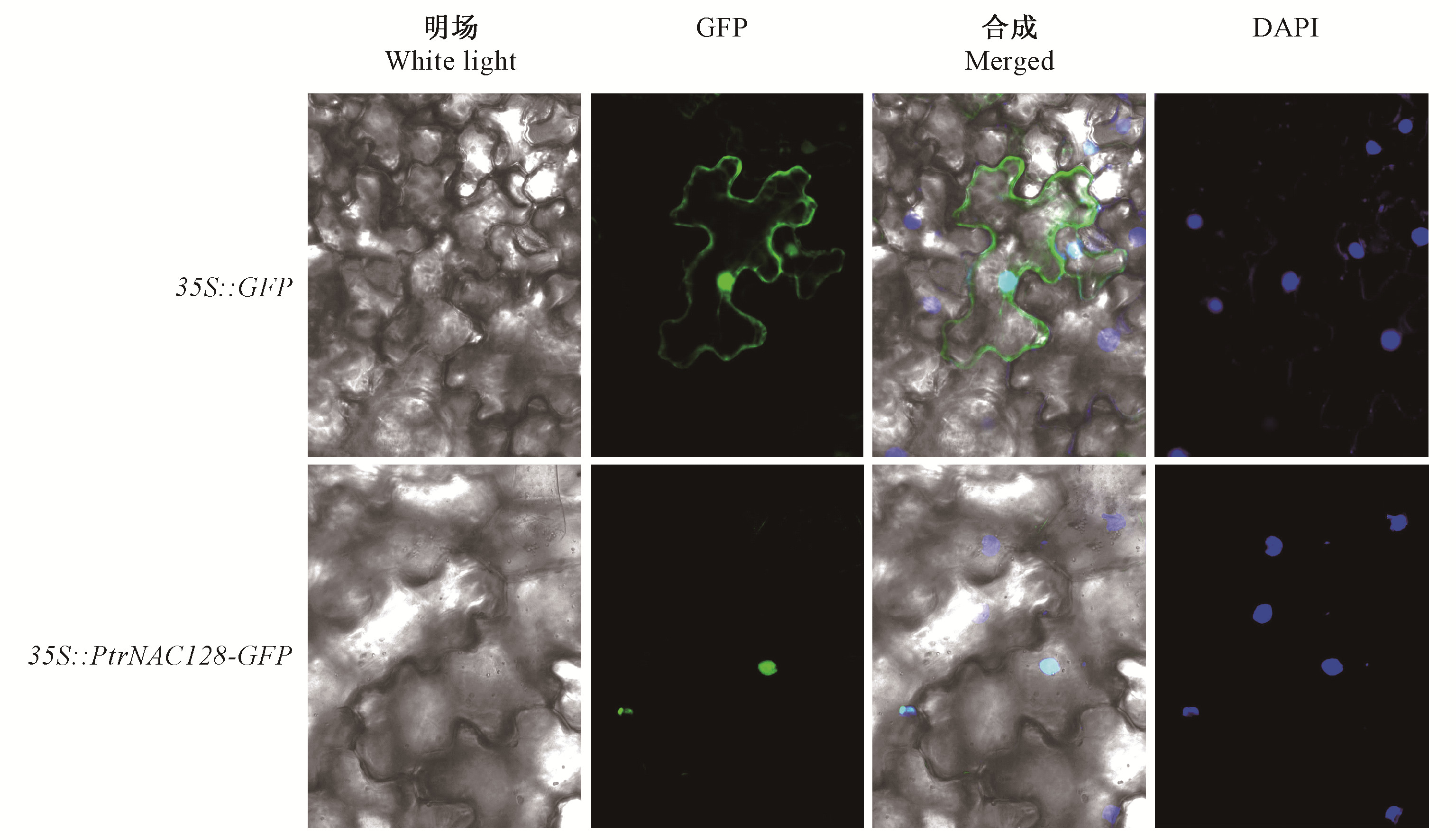
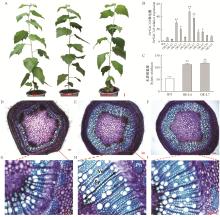
Fig.4
Effects of PtrNAC128 overexpression on secondary xylem of Populus tomentosa stems A. Phenotypes of representative 6-month-old wild-type(WT) and transgenic lines(OE-L6 and OE-L7); B. The relative expression levels of PtrNAC128 in transgenic(OE-L1-11) and wild-type poplar; C. Xylem width of wild-type and PtrNAC128 overexpression lines (OE-L6 and OE-L7). D, E, F are stem sections from the 6th internode of wild-type(D) and transgenic plants OE-L6(E) and OE-L7(F); G, H, I are the enlarged view of red rectangular in D, E, F, respectively. Ph: Phloem; Xy: Xylem; Ve: Vessel; Xf: Xylary fiber. The single and double stars indicate significant difference at 0.05 and 0.01 levels according to Student's t-test, the same below."

Table 2
Secondary cell wall composition analysis of the stems in PtrNAC128 overexpression and wild-type P. tomentosa"
| 次生壁组分 Secondary cell wall composition | 野生型 Wild type | 过表达PtrNAC128株系 PtrNAC128 overexpression plants | ||
| OE-L3 | OE-L6 | OE-L7 | ||
| 木质素Lignin | ||||
| 乙酰溴溶解AcBr-soluble/(mg·g-1) | 215.6±4.4 | 223.9±6.5* | 258.2±6.6* | 238.4±6.9* |
| Klason+酸溶Klason+acid-soluble/(mg·g-1) | 217.8±4.4 | 242.8±10.7 | 263.7±14.4* | 255.9±8.6* |
| Klason/(mg·g-1) | 189.7±2.2 | 212.7±5.8 | 231.9±4.1* | 225.4±4.3* |
| 酸溶Acid-soluble/(mg·g-1) | 28.1±2.2 | 30.1±4.0 | 31.8±4.2* | 30.5±4.1 |
| 半纤维素Hemicellulose/(mg·g-1) | 155.2±5.1 | 155.9±8.1 | 159.6±5.7 | 152.1±4.2 |
| 纤维素Cellulose/(mg·g-1) | 443.6±12.5 | 476.4±13.5* | 501.5±17.2* | 488.8±16.6* |
| 李慧, 郭晓蕊, 刘雅琳, 等. 木材形成过程中次生壁沉积和细胞程序性死亡的分子调控机制. 中国科学:生命科学, 2020, 50 (2): 123- 135. | |
| Li H , Guo X R , Liu Y L , et al. The molecular mechanism in secondary wall deposition and programmed cell death of wood formation. Scientia Sinica Vitae:Life Science, 2020, 50 (2): 123- 135. | |
| 孙利军, 李大勇, 张慧娟, 等. NAC转录因子在植物抗病和抗非生物胁迫反应中的作用. 遗传, 2012, 34 (8): 993- 1002. | |
| Sun L J , Li D Y , Zhang H J , et al. Functions of NAC transcription factors in biotic and abiotic stress responses in plants. Hereditas, 2012, 34 (8): 993- 1002. | |
| Aida M , Ishida T , Fukaki H , et al. Genes involved in organ separation in Arabidopsis: an analysis of the cup-shaped cotyledon mutant. Plant Cell, 1997, 9 (6): 841- 857. | |
| Dence C W . Methods in lignin chemistry. Berlin: Springer, 1992: 33- 61. | |
| Duval M , Hsieh T F , Kim S Y , et al. Molecular characterization of AtNAM:a member of the Arabidopsis NAC domain superfamily. Plant Mol Biol, 2002, 50 (2): 237- 248. | |
| Fujiwara S , Mitsuda N . ANAC075, a putative regulator of VASCULAR-RELATED NAC-DOMAIN7, is a repressor of flowering. Plant Biotechnology, 2016, 33 (4): 255- 265. | |
| Guo H S , Xie Q , Fei J F , et al. MicroRNA directs mRNA cleavage of the transcription factor NAC1 to downregulate auxin signals for Arabidopsis lateral root development. Plant Cell, 2005, 17 (5): 1376- 1386. | |
|
Hegedus D , Yu M , Baldwin D , et al. Molecular characterization of Brassica napus NAC domain transcriptional activators induced in response to biotic and abiotic stress. Plant Mol Biol, 2003, 53 (3): 383- 397.
doi: 10.1023/B:PLAN.0000006944.61384.11 |
|
|
Hu R , Qi G , Kong Y , et al. Comprehensive analysis of NAC domain transcription factor gene family in Populus trichocarpa. BMC Plant Biology, 2010, 10, 145.
doi: 10.1186/1471-2229-10-145 |
|
| Jensen M K , Kjaersgaard T , Nielsen M M , et al. The Arabidopsis thaliana NAC transcription factor family:structure-function relationships and determinants of ANAC019 stress signaling. Biochemical Journal, 2010, 426 (7): 183- 196. | |
| Jeong J S , Kim Y S , Redillas M C F , et al. OsNAC5 overexpression enlarges root diameter in rice plants leading to enhanced drought tolerance and increased grain yield in the field. Plant Biotechnology Journal, 2013, 11 (1): 101- 114. | |
|
Jung H J , Varel V H , Weimer P J , et al. Accuracy of Klason lignin and acid detergent lignin methods as assessed by bomb calorimetry. Journal of Agricultural and Food Chemistry, 1999, 47 (5): 2005- 2008.
doi: 10.1021/jf981250q |
|
|
Kim H J , Nam H G , Lim P O , et al. Regulatory network of NAC transcription factors in leaf senescence. Curr Opin Plant Biol, 2016, 33, 48- 56.
doi: 10.1016/j.pbi.2016.06.002 |
|
|
Lee C , Teng Q , Zhong R , et al. Molecular dissection of xylan biosynthesis during wood formation in poplar. Molecular Plant, 2011, 4 (4): 730- 747.
doi: 10.1093/mp/ssr035 |
|
|
Li C , Ma X , Yu H , et al. Ectopic expression of PtoMYB74 in poplar and Arabidopsis promotes secondary cell wall formation. Frontiers in Plant Science, 2018, 9, 1262.
doi: 10.3389/fpls.2018.01262 |
|
| Livak K J , Schmittgen T D . Analysis of relative gene expression data using real-time quantitative PCR. Methods, 2002, 25 (4): 402- 408. | |
|
Liyama K , Wallis A . An improved acetyl bromide procedure for determining lignin in woods and wood pulps. Wood Science & Technology, 1988, 22 (3): 271- 280.
doi: 10.1007/BF00386022 |
|
|
Mitsuda N , Iwase A , Yamamoto H , et al. NAC transcription factors, NST1 and NST3, are key regulators of the formation of secondary walls in woody tissues of Arabidopsis. Plant Cell, 2007, 19, 270- 280.
doi: 10.1105/tpc.106.047043 |
|
| Nakano Y , Yamaguchi M , Endo H , et al. NAC-MYB-based transcriptional regulation of secondary cell wall biosynthesis in land plants. Front Plant Sci, 2015, 6, 288. | |
|
Nuruzzaman M , Manimekalai R , Sharoni A M , et al. Genome-wide analysis of NAC transcription factor family in rice. Gene, 2010, 465 (1/2): 30- 44.
doi: 10.1186/1471-2229-10-145 |
|
| Ohashi-Ito K , Oda Y , Fukuda H , et al. Arabidopsis VASCULAR-RELATED NAC-DOMAIN6 directly regulates the genes that govern programmed cell death and secondary wall formation during xylem differentiation. Plant Cell, 2010, 22 (10): 3461- 3473. | |
| Ohtani M , Nishikubo N , Xu B , et al. A NAC domain protein family contributing to the regulation of wood formation in poplar. Plant Journal, 2011, 67 (3): 499- 512. | |
| Ooka H , Satoh K , Doi K , et al. Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res, 2003, 10 (6): 239- 247. | |
| Park J , Kim Y S , Kim S G , et al. Integration of auxin and salt signals by the NAC transcription factor NTM2 during seed germination in Arabidopsis. Plant Physiol, 2011, 156 (2): 537- 549. | |
| Sakamoto S , Mitsuda N . Reconstitution of a secondary cell wall in a secondary cell wall-deficient Arabidopsis mutant. Plant Cell Physiology, 2015, 56 (2): 299- 310. | |
| Shan X , Yang K , Xu X , et al. Genome-wide investigation of the NAC gene family and its potential association with the secondary cell wall in Moso bamboo. Biomolecules, 2019, 9 (10): 609. | |
| Shi R , Sun Y H , Li Q , et al. Towards a systems approach for lignin biosynthesis in Populus trichocarpa: transcript abundance and specificity of the monolignol biosynthetic genes. Plant Cell Physiology, 2010, 51 (1): 144- 163. | |
| Song D , Shen J , Li L . Characterization of cellulose synthase complexes in Populus xylem differentiation. New Phytologist, 2010, 187 (3): 777- 790. | |
| Sun H , Hu M L , Li J Y . Comprehensive analysis of NAC transcription factors uncovers their roles during fiber development and stress response in cotton. BMC Plant Biol, 2018, 8 (1): 150. | |
| Takata N , Awano T , Nakata M T , et al. Populus NST/SND orthologs are key regulators of secondary cell wall formation in wood fibers, phloem fibers and xylem ray parenchyma cells. Tree Physiology, 2019, 39 (4): 514- 525. | |
| Taylor-Teeples M , Lin L , de Lucas M , et al. An Arabidopsis gene regulatory network for secondary cell wall synthesis. Nature, 2015, 517, 571- 575. | |
| Tran L S , Nakashima K , Sakuma Y , et al. Isolation and functional analysis of Arabidopsis stress inducible NAC transcription factors that bind to a drought responsive cis-element in the early responsive to dehydration stress 1 promoter. Plant Cell, 2004, 16, 2481- 2498. | |
| Tuskan G A , Difazio S , Jansson S , et al. The genome of black cottonwood, Populus trichocarpa (Torr.& Gray). Science, 2006, 313 (5793): 1596- 1604. | |
| Van Soest P J , Wine R H . Use of detergents in the analysis of fibrous feeds IV:Determination of plant cell-wall constituents. Journal of the Association of Official Analytical Chemists, 1967, 50 (1): 50- 55. | |
| Wang T Z , Liu M , Zhao M G , et al. Identification and characterization of long non-coding RNAs involved in osmotic and salt stress in Medicago truncatula using genome-wide high-throughput sequencing. BMC Plant Biol, 2015, 15, 131. | |
| Xi W , Song D , Sun J , et al. Formation of wood secondary cell wall may involve two type cellulose synthase complexes in Populus. Plant Molecular Biology, 2016, 93 (4/5): 419- 429. | |
| Zhong R , Richardson E A , Ye Z H . The MYB46 transcription factor is a direct target of SND1 and regulates secondary wall biosynthesis in Arabidopsis. Plant Cell, 2007, 19 (9): 2776- 2792. | |
| Zhong R , Ye Z H . Transcriptional regulation of lignin biosynthesis. Plant Signaling & Behavior, 2009, 4 (11): 1028- 1034. | |
| Zhong R , Lee C , Ye Z H , et al. Evolutionary conservation of the transcriptional network regulating secondary cell wall biosynthesis. Trends Plant Sci, 2010, 15 (11): 625- 632. | |
| Zhong R , Lee C , McCarthy R L , et al. Transcriptional activation of secondary wall biosynthesis by rice and maize NAC and MYB transcription factors. Plant Cell Physiol, 2011a, 52 (10): 1856- 1871. | |
| Zhong R , McCarthy R L , Lee C , et al. Dissection of the transcriptional program regulating secondary wall biosynthesis during wood formation in poplar. Plant Physiol, 2011b, 157 (3): 1452- 1468. | |
| Zhong R , Ye Z H . Secondary cell walls:Biosynthesis, patterned deposition and transcriptional regulation. Plant Cell Physiol, 2014, 56, 195- 214. | |
| Zhong R , Ye Z H . The Arabidopsis NAC transcription factor NST2 functions together with SND1 and NST1 to regulate secondary wall biosynthesis in fibers of inflorescence stems. Plant Signaling & Behavior, 2015, 10 (2): e989746. |
| [1] | Weibo Sun,Xindong Gong,Yan Zhou,Hongyan Li. Photosynthetic Characteristics of Transgenic Poplars with Maize PEPC and PPDK Gene at Young Plant Stage [J]. Scientia Silvae Sinicae, 2020, 56(7): 33-43. |
| [2] | Xing Wu,Xingfeng Hu,Peizhen Chen,Xiaobo Sun,Fan Wu,Kongshu Ji. Cloning and Functional Analysis of PmPIN1 Gene from Pinus massoniana [J]. Scientia Silvae Sinicae, 2020, 56(3): 184-192. |
| [3] | Lei Zhang,Jianjun Hu. An Analysis of T-DNA Insertion Loci and Detection of the Locus-Specific of Transgenic Populus nigra Lines with BtCry1Ac [J]. Scientia Silvae Sinicae, 2020, 56(10): 45-52. |
| [4] | Weibo Sun,Zhaoqiong Wei,Xiaoxing Ma,Hui Wei,Qiang Zhuge. Safety Assessment of a Field Trial of Three Types of Transgenic Poplar Nanlin895 [J]. Scientia Silvae Sinicae, 2020, 56(10): 53-62. |
| [5] | Zhang Chao, Wang Jinmao, Zhao Jie, Pang Dingwei, Zhang Dejian, Yang Minsheng. Expression Characteristics of Bt Gene in Transgenic Poplar Transformed by Different Multi-Gene Vectors [J]. Scientia Silvae Sinicae, 2019, 55(9): 61-70. |
| [6] | Zhang Qin, Xu Zongda, Zhao Kai, Li Xiaowei, Zhang Luosha, Zhang Qixiang. Isolation and Biological Function Analysis of Anthocyanin Regulatory Gene PmMYB1 from Prunus mume [J]. Scientia Silvae Sinicae, 2018, 54(10): 64-72. |
| [7] | Jiang Wenhu, Zhang Dejian, Liu Junxia, Li Chaoli, Lu Zhanyuan, Yang Minsheng. Comparative Analysis of Arthropod Communities in Transgenic Bt and Non-Transgenic Poplar-Cotton Composite Systems [J]. Scientia Silvae Sinicae, 2018, 54(10): 73-79. |
| [8] | Chen Panfei, Zuo Lihui, Wang Guiying, Wang Jinmao, Ren Yachao, Yang Minsheng. Growth and Physiological Responses of Transgenic Populus×euramericana cv. ‘4/76’ with Multiple Genes Under Salt Stress [J]. Scientia Silvae Sinicae, 2017, 53(7): 45-53. |
| [9] | Huang Juan, Chen Cun, Zhang Weixi, Ding Changjun, Su Xiaohua, Huang Qinjun. Effects of Drought Stress on Anatomical Structure and Photosynthetic Characteristics of Transgenic JERF36 Populus alba×P. berolinensis Seedling Leaves [J]. Scientia Silvae Sinicae, 2017, 53(5): 8-15. |
| [10] | Yin Wu, Sun Weibo, Zhou Yan, Zhuge Qiang. Cloning and Functional Analysis of Rubisco Activase Gene from Populus trichocarpa [J]. Scientia Silvae Sinicae, 2017, 53(4): 83-95. |
| [11] | Chen Panfei, Ren Yachao, Zhang Jun, Wang Jinmao, Yang Minsheng. Expression and Transportation of Bt Toxic Protein in 8-Year-Old Grafted Transgenic Poplar [J]. Scientia Silvae Sinicae, 2016, 52(7): 46-52. |
| [12] | Zhu Wenxu, Zhang Bingyu, Huang Qinjun, Chu Yanguang, Ding Changjun, Zhang Weixi, Su Xiaohua. Effects of Multi-Gene Transgenic Populus×euramericana 'Guariento' on the Function of Microbial Population in the Rhizosphere Soil [J]. Scientia Silvae Sinicae, 2015, 51(11): 69-75. |
| [13] | Wang Lei, Su Hongyan, Gu Liang, Chang Zhiyuan, Chen Na. Screening and Verification of PtMPKs Interacting with PtMKK4 of Populus trichocarpa [J]. Scientia Silvae Sinicae, 2015, 51(10): 60-66. |
| [14] | Sun Haiwei;Liu Jing;Song Jian;Weng Manli;Luo Lei;Huang Yanyan;Zhang Hong;Niu Qinglin;Wang Bin;Feng Dianqi;. Molecular Identification and Cold-Resistance Analysis of the AmGS Transgenic Photinia×fraseri ‘Red Robin’ Plants [J]. Scientia Silvae Sinicae, 2012, 48(7): 30-38. |
| [15] | Yin Wu;Li Lisha;Wang Like;Wang Mingxiu;Zhuge Qiang. Analysis of Photosynthetic Characteristics of Transgenic Poplars with Maize PEPC Gene [J]. Scientia Silvae Sinicae, 2012, 48(6): 63-71. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||