

Scientia Silvae Sinicae ›› 2026, Vol. 62 ›› Issue (3): 1-12.doi: 10.11707/j.1001-7488.LYKX20250058
• Reviews • Previous Articles Next Articles
Jiayue Zhao1,Zhijie Zong1,Xiyuan Peng1,Zhiqiang Li1,Xingxia Ma2,*( )
)
Received:2025-02-05
Revised:2025-05-08
Online:2026-03-15
Published:2026-03-12
Contact:
Xingxia Ma
E-mail:mxxyln@139.com
CLC Number:
Jiayue Zhao,Zhijie Zong,Xiyuan Peng,Zhiqiang Li,Xingxia Ma. Research Progress on Enzymes and Metabolic Pathways Involved in Lignin Biodegradation by Fungi[J]. Scientia Silvae Sinicae, 2026, 62(3): 1-12.
Fig.2
Distribution of wood decay fungi across fungal lineages Highlighting the occurrence of white rot, brown rot, and soft rot fungi. White rot fungi (green) and brown rot fungi (orange) are primarily concentrated in the class Agaricomycetes of the phylum Basidiomycota, while soft rot fungi are mainly found in the phylum Ascomycota. Uncertain or rare distributions are not marked."
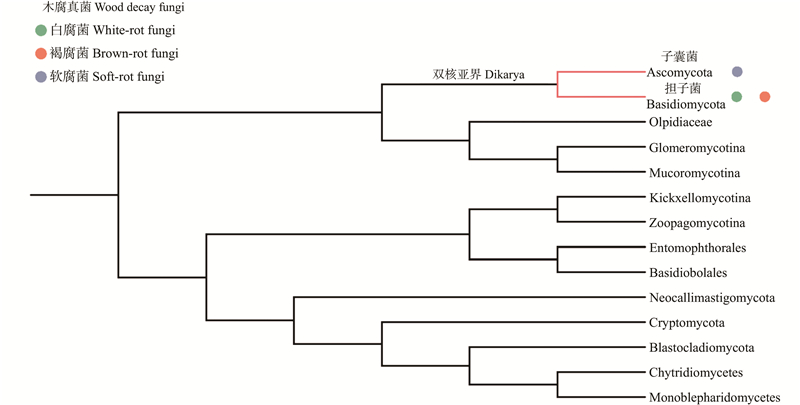
Table 1
Comparisons between c2092_g1_i1, lignin peroxidases, manganese peroxidases, and laccases"
| 特征Characteristics | c2092_g1_i1 | LiPs | MnPs | Laccases |
| 类型 Type | 氧化酶 Oxidase | 氧化酶 Oxidase | 氧化酶 Oxidase | 氧化酶 Oxidase |
| 分子量 Molecular weight | 44.5 kDa | 38~43 kDa | 38~62.5 kDa | 40~140 kDa 真菌漆酶Fungal laccase:40?140 kDa |
| 来源 Source | P. putredinis NO1在含小麦秸秆的 培养基的上清液 P. putredinis NO1 supernatant after incubation on wheat straw | 黄孢原毛平革菌 White-rot fungus Phanerochaete chrysosporium | 黄孢原毛平革菌 White-rot fungus Phanerochaete chrysosporium | 柬埔寨一种漆树科植物 A plant from Anacardiaceae in Cambodia |
| 蛋白家族 Protein family | 与中央酪氨酸酶结构域(PF00264) 同源,具有双核Ⅲ型铜结合位点 Homology to central tyrosinase domain (PF00264) with binuclear type-Ⅲ copper-binding site | Ⅱ类过氧化物酶 (PF00141) 含有血红素 ClassⅡ peroxidases (PF00141) with heme group | Ⅱ类过氧化物酶 (PF00141) 含有血红素和 Mn2+结合位点 ClassⅡ peroxidases (PF00141) with heme group and Mn2+ binding site | 多铜氧化酶 (PF00394) 含有Ⅰ、Ⅱ、Ⅲ型铜 结合位点 Multicopper oxidases (PF00394) with typeⅠ, Ⅱ, Ⅲ copper sites |
| 分布 Distribution | 在子囊菌纲类真菌中发现具 有>50%序列一致性的同源蛋白 Homologous proteins (>50 sequence identity) found in Sordariomycetes class of Ascomycetes | 存在于各种白腐菌中Found in various white-rot fungi | 存在于各种白腐菌中Found in various white-rot fungi | 广泛存在于真菌、植物和细菌中 Widely found in fungi, plants, and bacteria |
| 辅因子 Cofactors | 不依赖辅因子 Cofactor-independent | 一般不需要 Not typically required | Mn3?螯合和扩散 Required for Mn3+ chelation and diffusion | ABTS 或 HBT ABTS or HBT |
| 氧化特性 Oxidative nature | 厌氧环境下失活 Near-total loss of activity under anaerobic conditions | 厌氧条件下有活性 Active in anaerobic conditions | 厌氧条件下有活性 Active in anaerobic conditions | 厌氧条件下有活性 Active in anaerobic conditions but activity and mechanisms differ |
| 最佳pH值 Optimal pH | 10 | 根据来源和特定的 LiP 而异 Depend on the source and specific LiP | 根据来源和特定的 MnP 而异 Depend on the source and specific MnP | 根据来源和特定的 漆酶而异 Depend on the source and specific laccase |
| 最佳温度 Optimal temperature | 60 ℃ | 根据来源和特定的 LiP 而异 Depend on the source and specific LiP | 根据来源和特定的 MnP而异 Depend on the source and specific MnP | 根据来源和特定的 漆酶而异 Depend on the source and specific laccase |
| 木质素 降解特性 Lignin degradation | 具有β-醚酶活性、可从木质素中 释放麦黄酮 Effective tricin release and with β-etherase activity | 对酚类结构活性高 High for breaking non-phenolic bonds (β—O—4) | 对酚醛结构活性高 High for phenolic substrates, indirect oxidation via Mn3+ chelates | 对于酚键降解效果中等;介质参与效果增强 Moderate for phenolic bonds, enhanced by mediators |
| 潜在应用 Potential applications | 不依赖辅因子、适用于高温和碱性条件,可用于木质素改性和 木质纤维素降解 Cofactors-free, high-temperature and alkaline conditions, use in lignin modification and lignocellulose degradation | 降解木质素生产生物 燃料 Potential in lignin degradation for biofuel production | 与LiPs或laccases结合 Effective in combination with LiPs or laccases | 用途广泛,可用于生物修复、制浆漂白、生物传感 Versatile, used in bioremediation, pulp bleaching, and biosensors |
| 方 旋, 马星霞, 温敬伟, 等. 出土木质文物中分离真菌的生长特性研究. 北京林业大学学报, 2022, 44 (1): 123- 131. | |
| Fang X, Ma X X, Wen J W, et al. Growth characteristics of fungi isolated from unearthed wooden cultural relics. Journal of Beijing Forestry University, 2022, 44 (1): 123- 131. | |
|
李 坚, 甘文涛, 陈志俊, 等. 向新出发, 木材科学前沿发展. 森林工程, 2025, 41 (1): 1- 39.
doi: 10.7525/j.issn.1006-8023.2025.01.001 |
|
|
Li J, Gan W T, Chen Z J, et al. Frontier advances in wood science towards a new departure. Forest Engineering, 2025, 41 (1): 1- 39.
doi: 10.7525/j.issn.1006-8023.2025.01.001 |
|
|
方 旋, 张景朋, 李嘉欣, 等. 三种古建筑常用阔叶树材耐腐性及防腐可处理性研究. 木材科学与技术, 2024, 38 (2): 29- 35.
doi: 10.12326/j.2096-9694.2023145 |
|
|
Fang X, Zhang J M, Li J X, et al. The decay resistance and preservative treatability of three hardwoods commonly used in ancient buildings. Chinese Journal of Wood Science and Technology, 2024, 38 (2): 29- 35.
doi: 10.12326/j.2096-9694.2023145 |
|
| 马星霞, 蒋明亮, 李志强. 2011. 木材生物降解与保护. 北京: 中国林业出版社. | |
| Ma X X, Jiang M L, Li Z Q. 2011. Wood biodegradation and protection. Beijing: China Forestry Publishing House. [in Chinese] | |
| 孟 圆, 王逸凡, 李小雨, 等. 生物降解木质纤维素类生物质固废物的研究进展. 工业微生物, 2023, 53 (2): 43- 45. | |
| Meng Y, Wang Y F, Li X Y, et al. Research progress in biodegradation of lignocellulosic biomass solid waste. Industrial Microbiology, 2023, 53 (2): 43- 45. | |
|
熊怡心, 王 爽, 马星霞, 等. 木质素氧化酶系高产复合真菌培养体系构建. 林业科学, 2024, 60 (10): 133- 142.
doi: 10.11707/j.1001-7488.LYKX20230074 |
|
|
Xiong Y X, Wang S, Ma X X, et al. Construction of compound fungal culture system for high-yield ligninolytic enzymes. Scientia Silvae Sinicae, 2024, 60 (10): 133- 142.
doi: 10.11707/j.1001-7488.LYKX20230074 |
|
|
Abdel-Hamid A M, Solbiati J O, Cann I K O. Insights into lignin degradation and its potential industrial applications. Advances in Applied Microbiology, 2013, 82, 1- 28.
doi: 10.1016/b978-0-12-407679-2.00001-6 |
|
| Agosin E, Odier E. Solid-state fermentation, lignin degradation and resulting digestibility of wheat straw fermented by selected white-rot fungi. Applied Microbiology and Biotechnology, 1985, 21 (6): 397- 403. | |
|
Andlar M, Rezić T, Marđetko N, et al. Lignocellulose degradation: an overview of fungi and fungal enzymes involved in lignocellulose degradation. Engineering in Life Sciences, 2018, 18 (11): 768- 778.
doi: 10.1002/elsc.201800039 |
|
|
Atiwesh G, Parrish C C, Banoub J, et al. Lignin degradation by microorganisms: a review. Biotechnology Progress, 2022, 38 (2): e3226.
doi: 10.1002/btpr.3226 |
|
| Ayeronfe F, Kassim A, Ishak N, et al. A review on microbial degradation of lignin. Advanced Science Letters, 2018, 24 (6): 4407- 4413. | |
|
Baig K S. Interaction of enzymes with lignocellulosic materials: causes, mechanism and influencing factors. Bioresources and Bioprocessing, 2020, 7 (1): 21.
doi: 10.1186/s40643-020-00310-0 |
|
|
Billings A F, Fortney J L, Hazen T C, et al. Genome sequence and description of the anaerobic lignin-degrading bacterium Tolumonas lignolytica sp. nov. Standards in Genomic Sciences, 2015, 10 (1): 106.
doi: 10.1186/s40793-015-0100-3 |
|
|
Bomble Y J, Lin C Y, Amore A, et al. Lignocellulose deconstruction in the biosphere. Current Opinion in Chemical Biology, 2017, 41, 61- 70.
doi: 10.1016/j.cbpa.2017.10.013 |
|
|
Borneman W S, Hartley R D, Morrison W H, et al. Feruloyl and p-coumaroyl esterase from anaerobic fungi in relation to plant cell wall degradation. Applied Microbiology and Biotechnology, 1990, 33 (3): 345- 351.
doi: 10.1007/BF00164534 |
|
|
Chen J X, Zhang B Y, Luo L L, et al. A review on recycling techniques for bioethanol production from lignocellulosic biomass. Renewable and Sustainable Energy Reviews, 2021, 149, 111370.
doi: 10.1016/j.rser.2021.111370 |
|
|
Dashora K, Gattupalli M, Tripathi G D, et al. Fungal assisted valorisation of polymeric lignin: mechanism, enzymes and perspectives. Catalysts, 2023, 13 (1): 149.
doi: 10.3390/catal13010149 |
|
| Del Cerro C, Erickson E, Dong T, et al. 2021. Intracellular pathways for lignin catabolism in white-rot fungi. Proceedings of the National Academy of Sciences of the United States of America, 118(9): e2017381118. | |
| Ding S Y, Liu Y S, Zeng Y N, et al. 2012. How does plant cell wall nanoscale architecture correlate with enzymatic digestibility? Science, 338(6110): 1055−1060. | |
|
Duran K, Kohlstedt M, van Erven G, et al. From 13C-lignin to 13C-mycelium: Agaricus bisporus uses polymeric lignin as a carbon source. Science Advances, 2024, 10 (16): eadl3419.
doi: 10.1126/sciadv.adl3419 |
|
|
Duran K, Miebach J, van Erven G, et al. Oxidation-driven lignin removal by Agaricus bisporus from wheat straw-based compost at industrial scale. International Journal of Biological Macromolecules, 2023, 246, 125575.
doi: 10.1016/j.ijbiomac.2023.125575 |
|
|
Durrant A J, Wood D A, Cain R B. Lignocellulose biodegradation by Agaricus bisporus during solid substrate fermentation. Journal of General Microbiology, 1991, 137 (4): 751- 755.
doi: 10.1099/00221287-137-4-751 |
|
|
Fang X, Xiong Y X, Li J X, et al. Treatability changes of radiata pine heartwood induced by white-rot fungus Trametes versicolor. Forests, 2023, 14 (5): 1040.
doi: 10.3390/f14051040 |
|
|
Floudas D. Evolution of lignin decomposition systems in fungi. Advances in Botanical Research, 2021, 99, 37- 76.
doi: 10.1016/bs.abr.2021.05.003 |
|
|
Floudas D, Binder M, Riley R, et al. The Paleozoic origin of enzymatic lignin decomposition reconstructed from 31 fungal genomes. Science, 2012, 336 (6089): 1715- 1719.
doi: 10.1126/science.1221748 |
|
|
Fuchs G, Boll M, Heider J. Microbial degradation of aromatic compounds: from one strategy to four. Nature Reviews Microbiology, 2011, 9 (11): 803- 816.
doi: 10.1038/nrmicro2652 |
|
|
Gonzalez A, Corsini G, Lobos S, et al. Metabolic specialization and codon preference of lignocellulolytic genes in the white rot basidiomycete Ceriporiopsis subvermispora. Genes, 2020, 11 (10): 1227.
doi: 10.3390/genes11101227 |
|
|
Harwood C S, Parales R E. The beta-ketoadipate pathway and the biology of self-identity. Annual Review of Microbiology, 1996, 50 (1): 553- 590.
doi: 10.1146/annurev.micro.50.1.553 |
|
|
Hatakka A. Lignin-modifying enzymes from selected white-rot fungi: production and role from in lignin degradation. FEMS Microbiology Reviews, 1994, 13 (2/3): 125- 135.
doi: 10.1111/j.1574-6976.1994.tb00039.x |
|
|
Himmel M E, Ding S Y, Johnson D K, et al. Biomass recalcitrance: engineering plants and enzymes for biofuels production. Science, 2007, 315 (5813): 804- 807.
doi: 10.1126/science.1137016 |
|
|
Hofrichter M. Review: lignin conversion by manganese peroxidase (MnP). Enzyme and Microbial Technology, 2002, 30 (4): 454- 466.
doi: 10.1016/S0141-0229(01)00528-2 |
|
|
Hofrichter M, Vares T, Kalsi M, et al. Production of manganese peroxidase and organic acids and mineralization of 14C-labelled lignin (14C-DHP) during solid-state fermentation of wheat straw with the white rot fungus Nematoloma frowardii. Applied and Environmental Microbiology, 1999, 65 (5): 1864- 1870.
doi: 10.1128/AEM.65.5.1864-1870.1999 |
|
|
Iiyama K, Stone B A, MacAuley B J. Compositional changes in compost during composting and growth of Agaricus bisporus. Applied and Environmental Microbiology, 1994, 60 (5): 1538- 1546.
doi: 10.1128/aem.60.5.1538-1546.1994 |
|
|
Janusz G, Pawlik A, Sulej J, et al. Lignin degradation: microorganisms, enzymes involved, genomes analysis and evolution. FEMS Microbiology Reviews, 2017, 41 (6): 941- 962.
doi: 10.1093/femsre/fux049 |
|
|
Jurak E, Punt A M, Arts W, et al. Fate of carbohydrates and lignin during composting and mycelium growth of Agaricus bisporus on wheat straw based compost. PLoS One, 2015, 10 (10): e0138909.
doi: 10.1371/journal.pone.0138909 |
|
|
Kajikawa H, Kudo H, Kondo T, et al. Degradation of benzyl ether bonds of lignin by ruminal microbes. FEMS Microbiology Letters, 2000, 187 (1): 15- 20.
doi: 10.1111/j.1574-6968.2000.tb09129.x |
|
|
Kapich A, Hofrichter M, Vares T, et al. Coupling of manganese peroxidase-mediated lipid peroxidation with destruction of nonphenolic lignin model compounds and 14C-labeled lignins. Biochemical and Biophysical Research Communications, 1999, 259 (1): 212- 219.
doi: 10.1006/bbrc.1999.0742 |
|
|
Knežević A, Milovanović I, Stajić M, et al. Lignin degradation by selected fungal species. Bioresource Technology, 2013, 138, 117- 123.
doi: 10.1016/j.biortech.2013.03.182 |
|
|
Lankiewicz T S, Choudhary H, Gao Y, et al. Lignin deconstruction by anaerobic fungi. Nature Microbiology, 2023, 8, 596- 610.
doi: 10.1038/s41564-023-01336-8 |
|
|
Lindahl B D, Tunlid A. Ectomycorrhizal fungi - potential organic matter decomposers, yet not saprotrophs. New Phytologist, 2015, 205 (4): 1443- 1447.
doi: 10.1111/nph.13201 |
|
|
Liu Q Q, Luo L, Zheng L Q. Lignins: biosynthesis and biological functions in plants. International Journal of Molecular Sciences, 2018, 19 (2): 335.
doi: 10.3390/ijms19020335 |
|
|
Lubbers R J M, Dilokpimol A, Visser J, et al. A comparison between the homocyclic aromatic metabolic pathways from plant-derived compounds by bacteria and fungi. Biotechnology Advances, 2019, 37 (7): 107396.
doi: 10.1016/j.biotechadv.2019.05.002 |
|
| Lu Y, Lu Y C, Hu H Q, et al. Structural characterization of lignin and its degradation products with spectroscopic methods. Journal of Spectroscopy, 2017, 1, 8951658. | |
|
Ma F Y, Huang X, Ke M, et al. Role of selective fungal delignification in overcoming the saccharification recalcitrance of bamboo culms. ACS Sustainable Chemistry & Engineering, 2017a, 5 (10): 8884- 8894.
doi: 10.1021/acssuschemeng.7b01685 |
|
|
Ma X X, Jiang M L, Liu J L, et al. Preliminary analysis of amplicon high-throughput sequencing as a method for the assessment of fungal diversity in discolored wood. Holzforschung, 2017b, 71 (10): 793- 800.
doi: 10.1515/hf-2017-0015 |
|
|
Ma X X, Jiang M L, Wu Y Z, et al. Effect of wood surface treatment on fungal decay and termite resistance. BioResources, 2013, 8 (2): 2366- 2375.
doi: 10.15376/biores.8.2.2366-2375 |
|
| Masai E J, Katayama Y, Fukuda M. 2007. Genetic and biochemical investigations on bacterial catabolic pathways for lignin-derived aromatic compounds. Bioscience, Biotechnology, and Biochemistry, 71(1): 1−15. | |
| McLeod M P, Warren R L, Hsiao W W L, et al. The complete genome of Rhodococcus sp. RHA1 provides insights into a catabolic powerhouse. Proceedings of the National Academy of Sciences of the United States of America, 2006, 103 (42): 15582- 15587. | |
|
Middelhoven W J. Catabolism of benzene compounds by ascomycetous and basidiomycetous yeasts and yeastlike fungi: a literature review and an experimental approach. Antonie van Leeuwenhoek, 1993, 63 (2): 125- 144.
doi: 10.1007/BF00872388 |
|
|
Nakagame S, Chandra R P, Kadla J F, et al. Enhancing the enzymatic hydrolysis of lignocellulosic biomass by increasing the carboxylic acid content of the associated lignin. Biotechnology and Bioengineering, 2011, 108 (3): 538- 548.
doi: 10.1002/bit.22981 |
|
|
Nguyen T V T, Gye H, Baek H, et al. Selective adsorption of lignin peroxidase on lignin for biocatalytic conversion of poplar wood biomass to value-added chemicals. ACS Applied Materials & Interfaces, 2024, 16 (45): 62203- 62212.
doi: 10.1021/acsami.4c14971 |
|
| Oates N C, Abood A, Schirmacher A M, et al. 2021. A multi-omics approach to lignocellulolytic enzyme discovery reveals a new ligninase activity from Parascedosporium putredinis NO1. Proceedings of the National Academy of Sciences of the United States of America, 118(18): e2008888118. | |
| Pollegioni L, Tonin F, Rosini E. Lignin-degrading enzymes. The FEBS Journal, 2015, 282 (7): 1190- 1213. | |
|
Ragauskas A J, Beckham G T, Biddy M J, et al. Lignin valorization: improving lignin processing in the biorefinery. Science, 2014, 344 (6185): 1246843.
doi: 10.1126/science.1246843 |
|
|
Rahikainen J L, Evans J D, Mikander S, et al. Cellulase-lignin interactions-the role of carbohydrate-binding module and pH in non-productive binding. Enzyme and Microbial Technology, 2013, 53 (5): 315- 321.
doi: 10.1016/j.enzmictec.2013.07.003 |
|
|
Shao C, Ma X X, Jiang M L, et al. In-situ growth of hexaconazole/polydopamine/hexadecyltrimethoxysilane in multi-scale structured wood to prepare superhydrophobic wooden materials with decay resistance. Chemical Engineering Journal, 2023, 476, 146396.
doi: 10.1016/j.cej.2023.146396 |
|
|
Sammond D W, Yarbrough J M, Mansfield E, et al. Predicting enzyme adsorption to lignin films by calculating enzyme surface hydrophobicity. Journal of Biological Chemistry, 2014, 289 (30): 20960- 20969.
doi: 10.1074/jbc.M114.573642 |
|
| Sigoillot J C, Berrin J G, Bey M, et al. Fungal strategies for lignin degradation. Advances in Botanical Research, 2012, 61, 263- 308. | |
|
Silva J P, Ticona A R P, Hamann P R V, et al. Deconstruction of lignin: from enzymes to microorganisms. Molecules, 2021, 26 (8): 2299.
doi: 10.3390/molecules26082299 |
|
|
Sun S L, Huang Y, Sun R C, et al. The strong association of condensed phenolic moieties in isolated lignins with their inhibition of enzymatic hydrolysis. Green Chemistry, 2016, 18 (15): 4276- 4286.
doi: 10.1039/C6GC00685J |
|
|
Suryadi H, Judono J J, Putri M R, et al. Biodelignification of lignocellulose using ligninolytic enzymes from white-rot fungi. Heliyon, 2022, 8 (2): e08865.
doi: 10.1016/j.heliyon.2022.e08865 |
|
|
Susmel P, Stefanon B. Aspects of lignin degradation by rumen microorganisms. Journal of Biotechnology, 1993, 30 (1): 141- 148.
doi: 10.1016/0168-1656(93)90035-L |
|
|
Vane C H, Abbott G D, Head I M. The effect of fungal decay (Agaricus bisporus) on wheat straw lignin using pyrolysis-GC-MS in the presence of tetramethylammonium hydroxide (TMAH). Journal of Analytical and Applied Pyrolysis, 2001, 60 (1): 69- 78.
doi: 10.1016/S0165-2370(00)00156-X |
|
|
van Erven G, Kleijn A F, Patyshakuliyeva A, et al. Evidence for ligninolytic activity of the ascomycete fungus Podospora anserina. Biotechnology for Biofuels, 2020, 13, 1- 12.
doi: 10.1186/s13068-020-01713-z |
|
|
van Erven G, Nayan N, Sonnenberg A S M, et al. Mechanistic insight in the selective delignification of wheat straw by three white-rot fungal species through quantitative 13C-IS py-GC–MS and whole cell wall HSQC NMR. Biotechnology for Biofuels, 2018, 11, 262.
doi: 10.1186/s13068-018-1259-9 |
|
|
Weng C H, Peng X W, Han Y J. Depolymerization and conversion of lignin to value-added bioproducts by microbial and enzymatic catalysis. Biotechnology for Biofuels, 2021, 14 (1): 84.
doi: 10.1186/s13068-021-01934-w |
|
|
Yadav V K, Gupta N, Kumar P, et al. Recent advances in synthesis and degradation of lignin and lignin nanoparticles and their emerging applications in nanotechnology. Materials, 2022, 15 (3): 953.
doi: 10.3390/ma15030953 |
|
|
Yaguchi A, Franaszek N, O’Neill K, et al. Identification of oleaginous yeasts that metabolize aromatic compounds. Journal of Industrial Microbiology & Biotechnology, 2020, 47 (9/10): 801- 813.
doi: 10.1007/s10295-020-02269-5 |
|
|
Zhao J Y, Csetenyi L, Gadd G M. Biocorrosion of copper metal by Aspergillus niger. International Biodeterioration & Biodegradation, 2020, 154, 105081.
doi: 10.1016/j.ibiod.2020.105081 |
|
|
Zhao J Y, Csetenyi L, Gadd G M. Fungal-induced CaCO3 and SrCO3 precipitation: a potential strategy for bioprotection of concrete. Science of the Total Environment, 2022a, 816, 151501.
doi: 10.1016/j.scitotenv.2021.151501 |
|
|
Zhao J Y, Dyer T, Csetenyi L, et al. Fungal colonization and biomineralization for bioprotection of concrete. Journal of Cleaner Production, 2022b, 330, 129793.
doi: 10.1016/j.jclepro.2021.129793 |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||