

Scientia Silvae Sinicae ›› 2020, Vol. 56 ›› Issue (5): 69-79.doi: 10.11707/j.1001-7488.20200508
Special Issue: 林木育种
• Articles • Previous Articles Next Articles
Zhongyuan Liu,Zheng Liu,Ying Xu,Shanshan Liu,Zhilan Tian,Qingjun Xie,Caiqiu Gao*
Received:2019-04-01
Online:2020-05-25
Published:2020-06-13
Contact:
Caiqiu Gao
CLC Number:
Zhongyuan Liu,Zheng Liu,Ying Xu,Shanshan Liu,Zhilan Tian,Qingjun Xie,Caiqiu Gao. Cloning and Salt Tolerance Analysis of Transcription Factor HSFA4 from Betula platyphylla[J]. Scientia Silvae Sinicae, 2020, 56(5): 69-79.
Table 1
The primer sequences using in qRT-PCR and vector construction"
| 用途 Application | 引物名称 Primer name | 正向和反向引物 Forward and reverse primer (5′—3′) |
| qRT-PCR | BpHSFA4-F | AGGTTGCCAAGAATCGGTTACTTC |
| BpHSFA4-R | AGGCCGACTGTCAACATTAAGCTG | |
| Tubulin-F | TCAACCGCCTTGTCTCTCAGG | |
| Tubulin-R | TGGCTCGAATGCACTGTTGG | |
| Ubiquitin-F | GATTGAGGGGAGGGATGCTG | |
| Ubiquitin-R | GGAGGACAAGGTGGAGGGTG | |
| 载体构建 Vector construction | pROKⅡ-BpHSFA4-F | CGCTCTAGAATGGATGAAGCTCAGGGTGGTTCG |
| pROKⅡ-BpHSFA4-R | CGGGGTACCCTATCTTCCACTAAAATTTGCCAT | |
| pFGC5941-BpHSFA4-Cis-F | TTGGCGCGCCCTAACAGTAAAAACTCCAACGGAG | |
| pFGC5941-BpHSFA4-Cis-R | CGATTTAAATCCAGGGTTCTCAGTCAAGAACTGT | |
| pFGC5941-BpHSFA4-Anti-F | CCTCTAGACTAACAGTAAAAACTCCAACGGAG | |
| pFGC5941-BpHSFA4-Anti-R | TCGGATCCCCAGGGTTCTCAGTCAAGAACTGT |
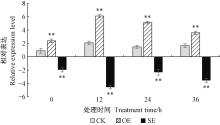
Fig.5
Expressions of BpHSFA4 in transient transgenic birch under 150 mmol ·L-1 NaCl stress CK:Plants transformed with empty pROKⅡ; OE:Plants transformed with overexpression of pROKⅡ-BpHSFA 4 ; SE:Plants transformed with RNAi-silencing of pFGC5941- BpHSFA4. Asterisks indicate Student's t-test: *, P < 0.05; **, P < 0.01. The same below. "
| 陈晓军, 叶春江, 吕慧颖, 等. GmHSFA1基因克隆及其过量表达提高转基因大豆的耐热性. 遗传, 2006. 28 (11): 1411- 1420. | |
| Chen X J , Ye C J , Lü H Y , et al. Cloning of GmHSFA1 gene and its overexpression leading to enhancement of heat tolerance in transgenic soybean. Hereditas, 2006. 28 (11): 1411- 1420. | |
| 胡涛, 张鸽香, 郑福超, 等. 植物盐胁迫响应的研究进展. 分子植物育种, 2018. 16 (9): 264- 273. | |
| Hu T , Zhang G X , Zheng F C , et al. Research progress in plant salt stress response. Molecular Plant Breeding, 2018. 16 (9): 264- 273. | |
| Bharti K , Schmidt E , Lyck R , et al. Isolation and characterization of HsfA3, a new heat stress transcription factor of Lycopersicon peruvianum. The Plant Journal, 2000. 22 (4): 355- 365. | |
| Chung E , Kim K M , Lee J H . Genome-wide analysis and molecular characterization of heat shock transcription factor family in Glycine max. Journal of Genetics and Genomics, 2013. 40 (3): 127- 135. | |
| Damberger F F , Pelton J G , Harrison C J , et al. Solution structure of the DNA-binding domain of the heat shock transcription factor determined by multidimensional heteronuclear magnetic resonance spectroscopy. Protein Science, 1994. 3 (10): 1806- 1821. | |
| Goswami S , Kumar R R , Sharma S K , et al. Calcium triggers protein kinases-induced signal transduction for augmenting the thermotolerance of developing wheat (Triticum aestivum) grain under the heat stress. Journal of Plant Biochemistry and Biotechnology, 2015. 24 (4): 441- 452. | |
| Guo J , Wu J , Ji Q , et al. Genome-wide analysis of heat shock transcription factor families in rice and Arabidopsis. Journal of Genetics and Genomics, 2008. 35 (2): 105- 118. | |
|
Guo M , Lu J P , Zhai Y F , et al. Genome-wide analysis, expression profile of heat shock factor gene family (CaHsfs) and characterisation of CaHsfA2 in pepper (Capsicum annuum L.).. BMC Plant Biology, 2015. 15 (1): 151.
doi: 10.1186/s12870-015-0512-7 |
|
| Hall T A . BioEdit:a user-friendly biological sequence alignment editor and analysis program for Windows95/98/NT. Nucleic Acids Symp Ser, 1999. 41, 95- 98. | |
| Hendry G A F , Baker A J M , Ewart C F . Cadmium tolerance and toxicity, oxygen radical processes and molecular damage in cadmium-tolerant and cadmium-sensitive clones of Holcus lanatus L. Acta Botanica Neerlandica, 1992. 41 (3): 271- 281. | |
|
Hossain M A , Bhattacharjee S , Armin S M , et al. Hydrogen peroxide priming modulates abiotic oxidative stress tolerance:insights from ROS detoxification and scavenging. Frontiers in Plant Science, 2015. 6, 420.
doi: 10.3389/fpls.2015.00420 |
|
| Huang X Y , Tao P , Li B Y , et al. Genome-wide identification, classification, and analysis of heat shock transcription factor family in Chinese cabbage (Brassica rapa pekinensis). Genetics Molecular Research, 2015. 14 (1): 2189- 2204. | |
| Jain G , Gould K S . Are betalain pigments the functional homologues of anthocyanins in plants. Environmental and Experimental Botany, 2015. 119, 48- 53. | |
| Kim M , Ahn J W , Jin U H , et al. Activation of the programmed cell death pathway by inhibition of proteasome function in plants. Journal of Biological Chemistry, 2003. 278, 19406- 19415. | |
| Li F , Zhang H , Zhao H , et al. Chrysanthemum CmHSFA4 gene positively regulates salt stress tolerance in transgenic chrysanthemum. Plant Biotechnology Journal, 2018. 16 (7): 1311- 1321. | |
|
Lin Y X , Jiang H Y , Chu Z X , et al. Genome-wide identification, classification and analysis of heat shock transcription factor family in maize. BMC Genomics, 2011. 12 (1): 76.
doi: 10.1186/1471-2164-12-76 |
|
| Liu Z , Wang P , Zhang T , et al. Comprehensive analysis of BpHSP genes and their expression under heat stresses in Betula platyphylla. Environmental and Experimental Botany, 2018. 152, 167- 176. | |
| Livak K J , Schittgen T D . Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods, 2001. 25 (4): 402- 408. | |
| Mishra S K . In the complex family of heat stress transcription factors, HsfA1 has a unique role as master regulator of thermotolerance in tomato. Genes Development, 2002. 16 (12): 1555- 1567. | |
| Mittler R . Abiotic stress, the field environment and stress combination. Trends in Plant Science, 2006. 11 (1): 15- 19. | |
| Perez-Salamo I , Papdi C , Rigo G , et al. The heat shock factor A4A confers salt tolerance and is regulated by oxidative stress and the mitogen-activated protein kinases MPK3 and MPK6. Plant Physiology, 2014. 165 (1): 319- 334. | |
|
Personat J M , Tejedor-Cano J , Prieto-Dapena P , et al. Co-overexpression of two Heat Shock Factors results in enhanced seed longevity and in synergistic effects on seedling tolerance to severe dehydration and oxidative stress. BMC Plant Biology, 2014. 14 (1): 56.
doi: 10.1186/1471-2229-14-56 |
|
| Qin F , Kakimoto M , Sakuma Y , et al. Regulation and functional analysis of ZmDREB2A in response to drought and heat stresses in Zea mays L. The Plant Journal, 2007. 50 (1): 54- 69. | |
| Rizhsky L , Liang H , Shuman J , et al. When defense pathways collide. The response of Arabidopsis to a combination of drought and heat stress. Plant Physiology, 2004. 134 (4): 1683- 1696. | |
| Scharf K D , Heider H , Höhfeld I , et al. The tomato Hsf system:HsfA2 needs interaction with HsfA1 for efficient nuclear import and may be localized in cytoplasmic heat stress granules. Molecular and Cellular Biology, 1998. 18 (4): 2240- 2251. | |
| Scharf K D , Rose S , Zott W , et al. Three tomato genes code for heat stress transcription factors with a region of remarkable homology to the DNA-binding domain of the yeast HSF. The EMBO Jurnal, 1990. 9 (13): 4495- 4501. | |
| Shim D , Hwang J U , Lee J , et al. Orthologs of the classA4 heat shock transcription factor HsfA4a confer cadmium tolerance in wheat and rice. The Plant Cell, 2009. 21 (12): 4031- 4043. | |
| Tamura K , Peterson D , Peterson N , et al. MEGA5:molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Molecular Biology and Evolution, 2011. 28 (10): 2731- 2739. | |
|
Wang J , Sun N , Deng T , et al. Genome-wide cloning, identification, classification and functional analysis of cotton heat shock transcription factors in cotton (Gossypium hirsutum). BMC Genomics, 2014. 15 (1): 961.
doi: 10.1186/1471-2164-15-961 |
|
|
Zhang J , Jia H , Li J , et al. Molecular evolution and expression divergence of the Populus euphratica Hsf genes provide insight into the stress acclimation of desert poplar. Scientific Reports, 2016. 6, 30050.
doi: 10.1038/srep30050 |
|
|
Zhang T , Zhao Y , Wang Y , et al. Comprehensive analysis of MYB gene family and their expressions under abiotic stresses and hormone treatments in Tamarix hispida. Frontiers in Plant Science, 2018. 9, 1303.
doi: 10.3389/fpls.2018.01303 |
|
| Zhang X , Wang L , Meng H , et al. Maize ABP9 enhances tolerance to multiple stresses in transgenic Arabidopsis by modulating ABA signaling and cellular levels of reactive oxygen species. Plant Molecular Biology, 2011. 75, 365- 378. | |
| Zhang Y , Wang Y , Wang C . Gene overexpression and gene silencing in birch using an Agrobacterium-mediated transient expression system. Molecular Biology Reports, 2012. 39, 5537- 5541. | |
| Zhou S , Zhang P , Jing Z , et al. Genome-wide identification and analysis of heat shock transcription factor family in cucumber (Cucumis sativus L.). Plant Omics:Journal of Plant Molecular Biology and Omics, 2013. 6 (6): 449- 455. |
| [1] | Zhang Chao, Wang Jinmao, Zhao Jie, Pang Dingwei, Zhang Dejian, Yang Minsheng. Expression Characteristics of Bt Gene in Transgenic Poplar Transformed by Different Multi-Gene Vectors [J]. Scientia Silvae Sinicae, 2019, 55(9): 61-70. |
| [2] | Liu Daofeng, Wang Xia, Dai Yin, Yang Jianfeng, Ma Jing, Li Mingyang, Sui Shunzhao. Cloning and Function Analysis of CpTAF10 from Wintersweet (Chimonanthus praecox) [J]. Scientia Silvae Sinicae, 2019, 55(6): 176-183. |
| [3] | Lu Huijun, Li Ziyi, Liang Hanyu, Yue Yuanzhi, Zhou Tianchang, Yang Yuzhang, Wang Yucheng, Ji Xiaoyu. Expression and Stress Tolerance Analysis of NAC24 from Tamarix hispida [J]. Scientia Silvae Sinicae, 2019, 55(3): 54-63. |
| [4] | Fan Weijian, Xiang Weifang, Wang Jing, Bai Penghua, Pan Lina, Yang Yixin, Zhu Gengping, Li Min. Effects of Insecticides and Ultraviolet on the Expression of sHSP12.2 Gene in Chouioia cunea [J]. Scientia Silvae Sinicae, 2019, 55(2): 128-136. |
| [5] | Yue Wang,Sufang Zhang,Yao Xu,Fu Liu,Xiangbo Kong,Zhen Zhang. Cloning and Expression Profiles of HcSID-1 Gene and Its Function Verification in Hyphantria cunea Larvae [J]. Scientia Silvae Sinicae, 2019, 55(10): 48-56. |
| [6] | Zhang Yang, Du Lin, Tang Xianfeng, Liu Huanhuan, Zhou Gongke, Chai Guohua. Identification and Expression Pattern Analyses of Populus BAG Genes [J]. Scientia Silvae Sinicae, 2019, 55(1): 138-145. |
| [7] | Wei Mingke, Yu Jinjian, Huang Xiaolong, Liu Qiongyao, Huang Huahong, Lin Erpei, Tong Zaikang. Cloning, Expression and Single Nucleotide Polymorphisms Analysis of NAC Transcription Factor Gene ClNAC1 in Cunninghamia lanceolata [J]. Scientia Silvae Sinicae, 2018, 54(9): 49-59. |
| [8] | Jin Mingyue, Jiang Feng, Jin Guangze, Liu Zhili. Variations of Specific Leaf Area in Different Growth Periods and Canopy Positions of Betula platyphylla at Different Ages [J]. Scientia Silvae Sinicae, 2018, 54(9): 18-26. |
| [9] | Wang Juan, Chen Hui. Identification and Expression of Cold Shock Domain-containing Protein Gene in Dendroctonus armandi [J]. Scientia Silvae Sinicae, 2018, 54(4): 58-66. |
| [10] | Zhang Hehua, Liu Jiaxin, Luo Chaobing, Zhang Lingyun. Cloning and Analysis of a Transcription Factor PwERF8 and the Promoter Sequences in Picea wilsonii [J]. Scientia Silvae Sinicae, 2018, 54(3): 48-60. |
| [11] | Li Jianbo, Jia Huixia, Zhang Jin, Liu Bobin, Hu Jianjun, Wang Lijuan, Lu Mengzhu. Effect of Overexpression of Populus tomentosa WUSCHEL-related homeobox 4 (PtoWOX4a) on the Secondary Growth of Poplar [J]. Scientia Silvae Sinicae, 2018, 54(2): 52-59. |
| [12] | Sun Huayu, Li Lichao, Zhao Hansheng, Yang Yihong, Wang Sining, Gao Zhimin. Molecular Characteristics and SSR Marker Development of Two NAC Genes from Daemonorops jenkinsiana [J]. Scientia Silvae Sinicae, 2017, 53(8): 132-140. |
| [13] | Tang Feifei, Zhao Yulin, Wang Peilong, Feng Deming, Song Yi, Gao Caiqiu. Cloning and Stress Tolerance Analysis of ThP5CR from Tamarix hispida [J]. Scientia Silvae Sinicae, 2017, 53(7): 1-9. |
| [14] | Chen Panfei, Zuo Lihui, Wang Guiying, Wang Jinmao, Ren Yachao, Yang Minsheng. Growth and Physiological Responses of Transgenic Populus×euramericana cv. ‘4/76’ with Multiple Genes Under Salt Stress [J]. Scientia Silvae Sinicae, 2017, 53(7): 45-53. |
| [15] | Li Nan, Zheng Yongqi, Ding Hongmei, Liu Xinhong, Sheng Weitong, Jiang Bo, Li Haibo. Analysis on Differential Expression of Cold Resistance Related Genes of Casuarina equisetifolia under Low Temperature Stress [J]. Scientia Silvae Sinicae, 2017, 53(7): 62-71. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||