

Scientia Silvae Sinicae ›› 2020, Vol. 56 ›› Issue (1): 38-53.doi: 10.11707/j.1001-7488.20200105
Previous Articles Next Articles
Xiaoyu Lu1,2,Zhu Chen1,Fei Tang1,Songling Fu2,Jie Ren1,*
Received:2019-02-01
Online:2020-01-25
Published:2020-02-24
Contact:
Jie Ren
CLC Number:
Xiaoyu Lu,Zhu Chen,Fei Tang,Songling Fu,Jie Ren. Combined Transcriptomic and Metabolomic Analysis Reveals Mechanism of Anthocyanin Changes in Red Maple(Acer rubrum) Leaves[J]. Scientia Silvae Sinicae, 2020, 56(1): 38-53.
Table 1
Name of anthocyanin derivatives"
| 代码Code | 中文名称Chinese name | 英文名称English name |
| Pg-H-M-C-R-G | 天竺葵素-O(羟基-甲氧基-鼠李糖基-吡喃葡糖苷)-O-吡喃葡糖苷 | Pelargonidin-O-(hydroxy-methoxy-cinnamoyl-L-rhamnopyranosyl-glucopyranoside)-O-glucopyranoside |
| Pg-G-H-G | 天竺葵素-O-(吡喃葡糖基-羟基-肉桂酸基-吡喃葡糖苷)-O-吡喃葡糖苷 | Pelargonidin-O-(glucopyranosyl-hydroxycinnamoyl-glucopyranoside)-O-glucopyranoside |
| Pg-Gsy | 天竺葵素-3-(2-谷氨基-葡糖基芸香糖苷) | Pelargonidin 3-(2-glu-glucosylrutinoside) |
| Pg-Ga | 天竺葵素-5-半乳糖苷 | Pelargonidin 5-galactoside |
| Cy-Acet | 矢车菊素-3-(4-乙酰葡糖苷) | Cyanidin 3-(4-acetylglucoside) |
| Cy-A-Ga | 矢车菊素-3-(6″-乙酰半乳糖苷) | Cyanidin 3-(6″-acetyl-galactoside) |
| Cy-X-H-G | 矢车菊素-O-(吡喃木糖基-羟基肉桂基-吡喃葡糖苷) | Cyanidin-O-(xylopyranosyl-hydroxycinnamoyl-glucopyranosyl) |
| Cy-S | 矢车菊素-3-桑布双糖苷 | Cyanidin 3-sambubioside |
| Cy-Arab | 矢车菊素-3-阿拉伯糖苷 | Cyanidin 3-arabinoside |
| Cy-L | 矢车菊素-3-昆布双糖苷 | Cyanidin 3-laminaribioside |
| Del-M-Glu | 飞燕草素-3-(6″-丙二葡糖苷) | Delphinidin 3-(6″-malonyl-glucoside) |
| Del-Acet | 飞燕草素-3-乙酰葡糖苷 | Delphinidin 3-acetylglucoside |
| Del-di-M | 飞燕草素-3, 5-二-(6-O-丙二葡糖苷) | Delphinidin 3, 5-di(6-O-malonylglucoside) |
| Del-S | 飞燕草素-3-桑布双糖苷 | Delphinidin 3-sambubioside |
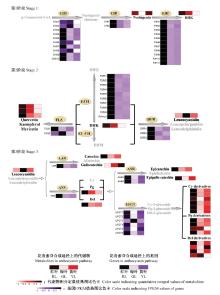
Fig.4
Detailed portion of the anthocyanin biosynthesis pathway that reveals the various expressions of related genes and different contents of metabolites CHS: Chalcone synthase; CHI: Chalcone isomerase; F3H: Flavanone 3-hydroxylase; F3′H: Flavanone 3′-hydroxylase; F3′5′H: Flavanone 3′, 5′-hydroxylase; DFR: Dihydroflavonol 4-reductase; ANS: Anthocyanidin synthase; UFGT: Flavonoid 3-O-glucosyltransferase; FLS: Flavonol synthase; LAR: Leucoanthocyanidin reductase; ANR: Anthocyanidin reductase; DHK: Dihydrokaempferol; DHQ: Dihydroquercetin; DHM: Dihydromyricetin; Cy: Cyanidin; Pg: Pelargonidin; Del: Delphinidin."
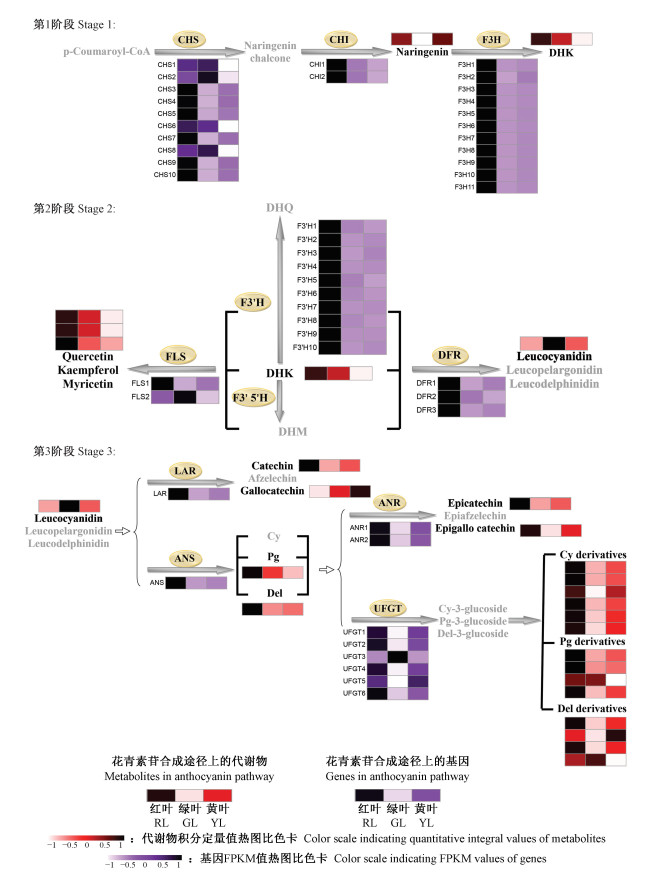
Table 2
Significantly enriched KEGG pathways in different comparison groups (corrected P≤0.01)"
| 编号No. | 通路名称Pathway | 通路ID Pathway ID | 通路注释的差异表达基因DEGs with pathway annotation | 通路注释的所有基因All genes with pathway annotation | P | 修正P值Corrected P |
| 红叶-绿叶Red leaves-green leaves | ||||||
| 1 | 核糖体Ribosome | ko03010 | 286 | 441 | 2.67E-23 | 3.26E-21 |
| 2 | 卟啉和叶绿素代谢Porphyrin and chlorophyll metabolism | ko00860 | 120 | 193 | 7.66E-10 | 4.67E-08 |
| 3 | 类黄酮生物合成Flavonoid biosynthesis | ko00941 | 76 | 102 | 4.37E-09 | 1.78E-07 |
| 4 | 光合作用-天线蛋白质Photosynthesis-antenna proteins | ko00196 | 58 | 76 | 1.58E-07 | 4.82E-06 |
| 5 | 精氨酸和脯氨酸代谢Arginine and proline metabolism | ko00330 | 171 | 377 | 5.79E-06 | 0.000 137 722 |
| 6 | 半胱氨酸和蛋氨酸代谢Cysteine and methionine metabolism | ko00270 | 224 | 525 | 6.77E-06 | 0.000 137 722 |
| 7 | 苯丙氨酸代谢Phenylalanine metabolism | ko00360 | 115 | 231 | 8.66E-06 | 0.000 150 948 |
| 8 | 精氨酸生物合成Arginine biosynthesis | ko00220 | 86 | 162 | 1.94E-05 | 0.000 295 595 |
| 9 | 甘氨酸、丝氨酸和苏氨酸代谢Glycine, serine and threonine metabolism | ko00260 | 142 | 312 | 2.97E-05 | 0.000 403 202 |
| 10 | 乙醛酸和二羧酸代谢Glyoxylate and dicarboxylate metabolism | ko00630 | 246 | 612 | 6.04E-05 | 0.000 736 680 |
| 11 | 抗坏血酸代谢Ascorbate and aldarate metabolism | ko00053 | 124 | 274 | 0.000 109 140 | 0.001 210 459 |
| 12 | 泛醌和其他萜类化合物-醌生物合成Ubiquinone and other terpenoid-quinone biosynthesis | ko00130 | 88 | 182 | 0.000 194 958 | 0.001 982 071 |
| 13 | 磷酸戊糖途径Pentose phosphate pathway | ko00030 | 151 | 360 | 0.000 349 958 | 0.003 284 225 |
| 14 | 苯丙氨酸、酪氨酸和色氨酸生物合成Phenylalanine, tyrosine and tryptophan biosynthesis | ko00400 | 87 | 186 | 0.000 479 922 | 0.004 182 176 |
| 15 | 植物-病原体相互作用Plant-pathogen interaction | ko04626 | 230 | 595 | 0.000 584 893 | 0.004 757 129 |
| 16 | 光合生物中的碳固定Carbon fixation in photosynthetic organisms | ko00710 | 229 | 593 | 0.000 625 676 | 0.004 770 781 |
| 17 | 谷胱甘肽代谢Glutathione metabolism | ko00480 | 158 | 388 | 0.000 727 198 | 0.005 094 233 |
| 18 | 苯丙烷类生物合成Phenylpropanoid biosynthesis | ko00940 | 96 | 214 | 0.000 751 608 | 0.005 094 233 |
| 19 | 氰氨基酸代谢Cyanoamino acid metabolism | ko00460 | 50 | 94 | 0.000 929 480 | 0.005 968 242 |
| 20 | 泛酸和CoA生物合成Pantothenate and CoA biosynthesis | ko00770 | 65 | 133 | 0.000 998 095 | 0.006 088 377 |
| 21 | 类胡萝卜素生物合成Carotenoid biosynthesis | ko00906 | 67 | 140 | 0.001 276 839 | 0.007 417 825 |
| 22 | 缬氨酸、亮氨酸和异亮氨酸的生物合成Valine, leucine and isoleucine biosynthesis | ko00290 | 55 | 110 | 0.001 581 266 | 0.008 768 837 |
| 23 | 果糖和甘露糖代谢Fructose and mannose metabolism | ko00051 | 125 | 304 | 0.001 850 311 | 0.009 814 695 |
| 24 | 二苯乙烯类、二芳基庚烷类和姜辣素类的生物合成Stilbenoid, diarylheptanoid and gingerol biosynthesis | ko00945 | 27 | 42 | 0.001 955 262 | 0.009 939 251 |
| 绿叶-黄叶Green leaves-yellow leaves | ||||||
| 1 | 核糖体Ribosome | ko03010 | 322 | 441 | 1.91E-13 | 2.33E-11 |
| 2 | 卟啉和叶绿素代谢Porphyrin and chlorophyll metabolism | ko00860 | 136 | 193 | 5.56E-06 | 0.000 339 316 |
| 红叶-黄叶Red leaves-yellow leaves | ||||||
| 1 | 类黄酮生物合成Flavonoid biosynthesis | ko00941 | 73 | 102 | 1.82E-10 | 2.22E-08 |
| 2 | 苯丙烷类生物合成Phenylpropanoid biosynthesis | ko00940 | 108 | 214 | 3.92E-08 | 2.39E-06 |
| 3 | 甘油脂代谢Glycerolipid metabolism | ko00561 | 202 | 520 | 1.16E-06 | 4.73E-05 |
| 4 | 调节自噬Regulation of autophagy | ko04140 | 118 | 287 | 2.67E-05 | 0.000 812 852 |
| 5 | 核糖体Ribosome | ko03010 | 164 | 441 | 6.85E-05 | 0.001 671 012 |
| 6 | 淀粉和蔗糖代谢Starch and sucrose metabolism | ko00500 | 370 | 1 149 | 0.000 168 590 | 0.003 427 987 |
| 7 | 苯丙氨酸、酪氨酸和色氨酸生物合成Phenylalanine, tyrosine and tryptophan biosynthesis | ko00400 | 78 | 186 | 0.000 350 292 | 0.005 966 480 |
| 8 | 糖胺聚糖降解Glycosaminoglycan degradation | ko00531 | 23 | 33 | 0.000 391 245 | 0.005 966 480 |
| 9 | N-聚糖生物合成N-glycan biosynthesis | ko00510 | 75 | 182 | 0.000 664 722 | 0.009 010 676 |
| 10 | 果糖和甘露糖代谢Fructose and mannose metabolism | ko00051 | 113 | 304 | 0.000 842 426 | 0.009 555 408 |
| 11 | 萜类骨架生物合成Terpenoid backbone biosynthesis | ko00900 | 67 | 160 | 0.000 896 156 | 0.009 555 408 |
| 12 | 脂肪酸延伸率Fatty acid elongation | ko00062 | 25 | 41 | 0.000 939 876 | 0.009 555 408 |
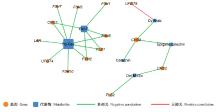
Fig.6
Strong correlations between anthocyanin biosynthesis-related genes and anthocyanin-related metabolites in red maple CHS: Chalcone synthase encoding gene; F3H: Flavanone 3-hydroxylase encoding gene; F3′H: Flavanone 3′-hydroxylase encoding gene; UFGT: Flavonoid 3-O-glucosyltransferase encoding gene; LAR: Leucoanthocyanidin reductase encoding gene; FLS: Flavonol synthase encoding gene. The metabolites Pg-Gsy, Del-S, Del-M-Glu, Cy-Arab are shown in Tab. 1."
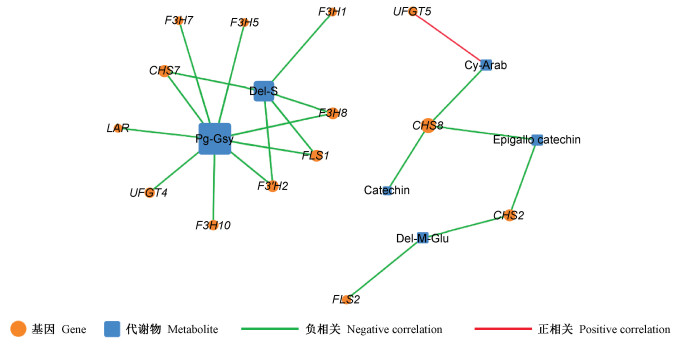
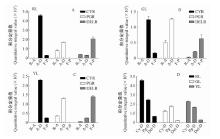
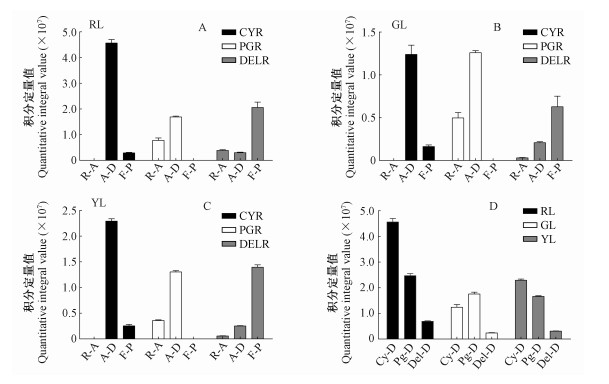
Fig.7
Content of anthocyanidins, anthocyanidin derivatives, and flavonoid products in red leaves(A), green leaves(B), yellow leaves(C) and content of three types of anthocyanidin-related pigments in red, green, and yellow leaves(D) RL: Red leaves; GL: Green leaves; YL: Yellow leaves; CYR: Cyanidin reactions; PGR: Pelargonidin reactions; DELR: Delphinidin reactions; R-A: Residual anthocyanidin; A-D: Anthocyanidin derivatives; F-P: Flavonoid product; Cy-D: Cyanidin derivatives; Pg-D: Pelargonidin derivatives; Del-D: Delphinidin derivatives."
| 冯立娟, 苑兆和, 尹燕雷, 等. 美国红枫叶色表达期间相关物质的研究. 山东林业科, 2008, (4): 9- 11. | |
| Feng L J , Yuan Z H , Yin Y L , et al. Studies on the related substances for color expression in American maple during the leaf color transition. Journal of Shandong Forestry Science & Technology, 2008, (4): 9- 11. | |
| 李玲. 2013. 五种红色叶植物叶片色素分析. 西安: 西北农林科技大学硕士学位论文. | |
| Li L . The research on the chromogenic pigment of leaves in five red plants. Xi'an: MS thesis of Northwest A&F University, 2013. | |
| 任杰, 丁增成, 唐菲, 等. 加拿大红枫的引种及繁育技术研究. 中国农学通报, 2013, 29 (1): 37- 41. | |
| Ren J , Ding Z C , Tang F , et al. Study on introduction and propagation techniques of red maple(Acer rubrum L. ). Chinese Agricultural Science Bulletin, 2013, 29 (1): 37- 41. | |
| 石柏林, 吴家森, 钟泰林. 6种槭树属植物种子特性及其发芽试验. 浙江林业科技, 2006, 26 (3): 38- 40. | |
| Shi B L , Wu J S , Zhong T L . Research on seed properties and germination test of six species of Acer. Journal of Zhejiang Forestry Science & Technology, 2006, 26 (3): 38- 40. | |
| 夏涛, 高丽萍. 类黄酮及茶儿茶素生物合成途径及其调控研究进展. 中国农业科学, 2009, 42 (8): 2899- 2908. | |
| Xia T , Gao L P . Advances in biosynthesis pathways and regulation of flavonoids and catechins. Scientia Agricultura Sinica, 2009, 42 (8): 2899- 2908. | |
| Abouzaid M M , Helson B V , Nozzolillo C , et al. Ethyl m-digallate from red maple, Acer rubrum L. , as the major resistance factor to forest tent caterpillar, Malacosoma disstria Hbn. Journal of Chemical Ecology, 2001, 27 (12): 2517- 2527. | |
|
Abrams M D . The red maple paradox. Bioscience, 1998, 48 (5): 355- 364.
doi: 10.2307/1313374 |
|
|
Alexander H D , Arthur M A . Implications of a predicted shift from upland oaks to red maple on forest hydrology and nutrient availability. Canadian Journal of Forest Research, 2010, 40 (4): 716- 726.
doi: 10.1139/X10-029 |
|
|
Cai Y , Weng K , Guo Y , et al. An integrated targeted metabolomic platform for high-throughput metabolite profiling and automated data processing. Metabolomics, 2015, 11 (6): 1575- 1586.
doi: 10.1007/s11306-015-0809-4 |
|
|
Chen Z , Lu X Y , Xuan Y , et al. Transcriptome analysis based on a combination of sequencing platforms provides insights into leaf pigmentation in Acer rubrum. BMC Plant Biology, 2019, 19 (1): 240- 256.
doi: 10.1186/s12870-019-1850-7 |
|
|
Cho K , Cho K S , Sohn H B , et al. Network analysis of the metabolome and transcriptome reveals novel regulation of potato pigmentation. Journal of Experimental Botany, 2016, 67 (5): 1519.
doi: 10.1093/jxb/erv549 |
|
| Chu Y X , Chen H R , Wu A Z , et al. Expression analysis of dihydroflavonol 4-reductase genes in Petunia hybrida. Genetics & Molecular Research, 2015, 14 (2): 5010- 5021. | |
|
Cole T , Williams B A , Geo P , et al. Transcript assembly and abundance estimation from RNA-Seq reveals thousands of new transcripts and switching among isoforms. Nature Biotechnology, 2010, 28 (5): 511- 515.
doi: 10.1038/nbt.1621 |
|
|
Davies K M , Schwinn K E , Deroles S C , et al. Enhancing anthocyanin production by altering competition for substrate between flavonol synthase and dihydroflavonol 4-reductase. Euphytica, 2003, 131 (3): 259- 268.
doi: 10.1023/A:1024018729349 |
|
|
Deng X , Bashandy H , Ainasoja M , et al. Functional diversification of duplicated chalcone synthase genes in anthocyanin biosynthesis of Gerbera hybrida. New Phytologist, 2014, 201 (4): 1469- 1483.
doi: 10.1111/nph.12610 |
|
| Deshmukh R , Sonah H , Patil G , et al. Integrating omic approaches for abiotic stress tolerance in soybean. Frontiers in Plant Science, 2014, 5, 244. | |
|
Dunn W B , Broadhurst D , Begley P , et al. Procedures for large-scale metabolic profiling of serum and plasma using gas chromatography and liquid chromatography coupled to mass spectrometry. Nature Protocols, 2011, 6 (7): 1060- 1083.
doi: 10.1038/nprot.2011.335 |
|
|
Ford C M , Boss P K , Hoj P B . Cloning and characterization of Vitis vinifera UDP-glucose: Flavonoid 3-O-glucosyltransferase, a homologue of the enzyme encoded by the maize Bronze-1 locus that may primarily serve to glucosylate anthocyanidins in vivo. Journal of Biological Chemistry, 1998, 273 (15): 9224- 9233.
doi: 10.1074/jbc.273.15.9224 |
|
|
Geoffroy T R , Fortin Y , Stevanovic T . Process optimisation for pilot-scale production of maple bark extracts, natural sources of antioxidants, phenolics, and carbohydrates. Chemical Papers, 2018, 72 (5): 1125- 1137.
doi: 10.1007/s11696-017-0355-9 |
|
|
Guo X , Chao Y , Le L , et al. Transcriptome of the floral transition in Rosa chinensis 'Old Blush'. Bmc Genomics, 2017, 18, 199.
doi: 10.1186/s12864-017-3584-y |
|
|
Han Y , Huang K , Liu Y , et al. Functional analysis of two flavanone-3-hydroxylase genes from Camellia sinensis: A critical role in flavonoid accumulation. Genes, 2017, 8 (11): 300.
doi: 10.3390/genes8110300 |
|
|
Hauke J , Kossowski T . Comparison of values of Pearson's and Spearman's correlation coefficients on the same sets of data. Quaestiones Geographicae, 2011, 30 (2): 87- 93.
doi: 10.2478/v10117-011-0021-1 |
|
|
He H , Ke H , Keting H , et al. Flower colour modification of chrysanthemum by suppression of F3'H and overexpression of the exogenous Senecio cruentus F3'5'H gene. PLoS One, 2013, 8 (11): e74395.
doi: 10.1371/journal.pone.0074395 |
|
| He J , Giusti M M . Anthocyanins: natural colorants with health-promoting properties. Annual Review of Food Science & Technology, 2010, 1 (1): 163- 187. | |
|
Hu C , Gong Y , Jin S , et al. Molecular analysis of a UDP-glucose: flavonoid 3-O-glucosyltransferase(UFGT) gene from purple potato (Solanum tuberosum). Molecular Biology Reports, 2011, 38 (1): 561- 567.
doi: 10.1007/s11033-010-0141-z |
|
| Kalubi K N , Mehes-Smith M , Omri A . Comparative analysis of metal translocation in red maple (Acer rubrum) and trembling aspen(Populus tremuloides) populations from stressed ecosystems contaminated with metals. Chemistry & Ecology, 2016, 32 (4): 312- 323. | |
| Kitada C , Gong Z , Tanaka Y , et al. Differential expression of two cytochrome P450s involved in the biosynthesis of flavones and anthocyanins in chemo-varietal forms of Perilla frutescens. Plant & Cell Physiology, 2001, 42 (12): 1338- 1344. | |
|
Kobayashi S , Ishimaru M , Ding C K , et al. Comparison of UDP-glucose: flavonoid 3-O-glucosyltransferase (UFGT) gene sequences between white grapes (Vitis vinifera) and their sports with red skin. Plant Science, 2001, 160 (3): 543- 550.
doi: 10.1016/S0168-9452(00)00425-8 |
|
|
Koes R , Verweij W , Quattrocchio F . Flavonoids: a colorful model for the regulation and evolution of biochemical pathways. Trends in Plant Science, 2005, 10 (5): 236- 242.
doi: 10.1016/j.tplants.2005.03.002 |
|
|
Koes R E , Spelt C E , Mol J N M . The chalcone synthase multigene family of Petunia hybrida (V30): differential, light-regulated expression during flower development and UV light induction. Plant Molecular Biology, 1989, 12 (2): 213- 225.
doi: 10.1007/BF00020506 |
|
| Kuittinen H , Aguadé M . Nucleotide variation at the chalcone isomerase locus in Arabidopsis thaliana. Genetics, 2000, 155 (2): 863- 872. | |
|
Kumar A , Singh B , Singh K . Functional characterization of flavanone 3-hydroxylase gene from Phyllanthus emblica (L.). Journal of Plant Biochemistry & Biotechnology, 2015, 24 (4): 453- 460.
doi: 10.1007/s13562-014-0296-0 |
|
|
Li B , Dewey C N . RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. Bmc Bioinformatics, 2011, 12 (1): 323- 323.
doi: 10.1186/1471-2105-12-323 |
|
| Li L , Liu Y , Liu Y , et al. Physiological response and resistance of three cultivars of Acer rubrum L. to continuous drought stress. Acta Ecologica Sinica, 2015, 35 (6): 196- 202. | |
|
Liu Y , Shi Z , Maximova S , et al. Proanthocyanidin synthesis in Theobroma cacao: genes encoding anthocyanidin synthase, anthocyanidin reductase, and leucoanthocyanidin reductase. BMC Plant Biology, 2013, 13 (1): 202.
doi: 10.1186/1471-2229-13-202 |
|
|
Lu X , Chen Z , Gao J , et al. Combined metabolome and transcriptome analyses of photosynthetic pigments in red maple. Plant Physiol Biochem, 2020, 154, 476- 490.
doi: 10.1016/j.plaphy.2020.06.025 |
|
|
Matile P , Hortensteiner S , Thomas H . Chlorophyll degradation. Annual Review of Plant Biology, 2006, 57, 55- 77.
doi: 10.1146/annurev.arplant.57.032905.105212 |
|
|
Moreno-Risueno M A , Busch W , Benfey P N . Omics meet networks—using systems approaches to infer regulatory networks in plants. Current Opinion in Plant Biology, 2010, 13 (2): 126- 131.
doi: 10.1016/j.pbi.2009.11.005 |
|
|
Moschen S , Bengoa Luoni S , Di Rienzo J A , et al. Integrating transcriptomic and metabolomic analysis to understand natural leaf senescence in sunflower. Plant Biotechnol J, 2016, 14 (2): 719- 734.
doi: 10.1111/pbi.12422 |
|
| Nam-Soo K , Min-Ji I , Kabwe N . Determination of DNA methylation associated with Acer rubrum(red maple) adaptation to metals: analysis of global DNA modifications and methylation-sensitive amplified polymorphism. Ecology & Evolution, 2016, 6 (16): 5749- 5760. | |
| Nkongolo K , Theriault G , Michael P . Differential levels of gene expression and molecular mechanisms between red maple (Acer rubrum) genotypes resistant and susceptible to nickel toxicity revealed by transcriptome analysis. Ecology & Evolution, 2018, 8 (10): 4876- 4890. | |
|
Piero A R L , Consoli A , Puglisi I , et al. Anthocyaninless cultivars of sweet orange lack to express the UDP-glucose flavonoid 3-O-glucosyl transferase. Journal of Plant Biochemistry & Biotechnology, 2005, 14 (1): 9- 14.
doi: 10.1007/BF03263217 |
|
|
Qian L , Liu Y , Qi Y , et al. Transcriptome sequencing and metabolite analysis reveals the role of delphinidin metabolism in flower colour in grape hyacinth. Journal of Experimental Botany, 2014, 65 (12): 3157- 3164.
doi: 10.1093/jxb/eru168 |
|
|
Rai A , Saito K , Yamazaki M . Integrated omics analysis of specialized metabolism in medicinal plants. Plant Journal, 2017, 90 (4): 764- 787.
doi: 10.1111/tpj.13485 |
|
|
Rai A , Yamazaki M , Takahashi H , et al. RNA-seq transcriptome analysis of Panax japonicus, and its comparison with other Panax species to identify potential genes involved in the saponins biosynthesis. Frontiers in Plant Science, 2016, 7, 148.
doi: 10.3389/fpls.2016.00481 |
|
|
Rebeki A , Lonari Z , Petrovi S , et al. Pearson's or Spearman's correlation coefficient: Which one to use?. Poljoprivreda, 2015, 21 (2): 47- 54.
doi: 10.18047/poljo.21.2.8 |
|
|
Riañopachón D M , Nagel A , Neigenfind J , et al. The GABI primary database: GABIPD—integration of plant 'omics'-data in gene context. Nature Precedings, 2009,
doi: 10.1038/npre.2008.1684.1 |
|
| Royer M , Diouf P N , Stevanovic T . Polyphenol contents and radical scavenging capacities of red maple (Acer rubrum L. ) extracts. Food & Chemical Toxicology, 2011, 49 (9): 2180- 2188. | |
|
Sapna K , Nie J , Chen H S , et al. Evaluation of gene association methods for coexpression network construction and biological knowledge discovery. PLoS ONE, 2012, 7 (11): e50411.
doi: 10.1371/journal.pone.0050411 |
|
|
Schmitzer V , Osterc G , Veberic R , et al. Correlation between chromaticity values and major anthocyanins in seven Acer palmatum Thunb. cultivars. Scientia Horticulturae, 2009, 119 (4): 442- 446.
doi: 10.1016/j.scienta.2008.09.003 |
|
|
Shen Q , Fu L , Dai F , et al. Multi-omics analysis reveals molecular mechanisms of shoot adaption to salt stress in Tibetan wild barley. Bmc Genomics, 2016, 17, 889.
doi: 10.1186/s12864-016-3242-9 |
|
|
Shimada N , Ayabe S I . A cluster of genes encodes the two types of chalcone isomerase involved in the biosynthesis of general flavonoids and legume-specific 5-deoxy(iso)flavonoids in Lotus japonicus. Plant Physiology, 2003, 131 (3): 941- 951.
doi: 10.1104/pp.004820 |
|
| Sibley J L , Eakes D J , Gilliam C H , et al. Foliage characteristics of selected red maple cultivars. Research Report, 1995, 1 (1): 40- 41. | |
|
Smith C A , Want E J , O'maille G , et al. XCMS: Processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Analytical Chemistry, 2006, 78 (3): 779- 787.
doi: 10.1021/ac051437y |
|
|
Song T T , Xu H H , Sun N , et al. Metabolomic analysis of alfalfa (Medicago sativa L. ) root-symbiotic rhizobia responses under alkali stress.. Frontiers in Plant Science, 2017, 8, 1208.
doi: 10.3389/fpls.2017.0120 |
|
|
Springob K , Nakajima J I , Yamazaki M , et al. Recent advances in the biosynthesis and accumulation of anthocyanins. Natural Product Reports, 2003, 20 (3): 288- 303.
doi: 10.1039/b109542k |
|
|
Tanaka Y , Sasaki N , Ohmiya A . Biosynthesis of plant pigments: anthocyanins, betalains and carotenoids. Plant Journal, 2008, 54 (4): 733- 749.
doi: 10.1111/j.1365-313X.2008.03447.x |
|
|
Wang J , Zhang T , Shen X , et al. Serum metabolomics for early diagnosis of esophageal squamous cell carcinoma by UHPLC-QTOF/MS. Metabolomics, 2016, 12 (7): 1- 10.
doi: 10.1007/s11306-016-1050-5 |
|
|
Want E J , Wilson I D , Gika H , et al. Global metabolic profiling procedures for urine using UPLC-MS. Nature Protocols, 2010, 5 (6): 1005- 1018.
doi: 10.1038/nprot.2010.50 |
|
|
Williams R S , Lincoln D E , Norby R J . Development of gypsy moth larvae feeding on red maple saplings at elevated CO2 and temperature. Oecologia, 2003, 137 (1): 114- 122.
doi: 10.1007/s00442-003-1327-z |
|
| Wu X , Prior R L . Systematic identification and characterization of anthocyanins by HPLC-ESI-MS/MS in common foods in the United States: fruits and berries. Journal of Agricultural & Food Chemistry, 2005, 53 (7): 2589- 2599. | |
| Zheng Y , Wang C Y , Wang S Y , et al. Effect of high-oxygen atmospheres on blueberry phenolics, anthocyanins, and antioxidant capacity. Journal of Agricultural & Food Chemistry, 2003, 51 (24): 7162- 7169. |
| [1] | Peihuang Zhu,Yu Chen,Lingzhi Zhu,Rong Li,Kongshu Ji. Codon Usage Bias and Its Influencing Factors in Pinus massoniana Transcriptome [J]. Scientia Silvae Sinicae, 2020, 56(4): 74-81. |
| [2] | Zhao Qingquan, Chi Yujie, Zhang Jian, Feng Lianrong. Transcriptome Construction and Related Gene Expression Analysis of Lenzites gibbosa in Woody Environment [J]. Scientia Silvae Sinicae, 2019, 55(8): 95-105. |
| [3] | Han Xiaohong, Lu Ciding, Hua Yin, Lin Haoyu, Shi Yufei, Wu Songqing, Zhang Feiping, Liang Guanghong. Phylogenetic Analysis of Transcriptome and Three Detoxification Enzyme Families Related Genes in Anoplophora chinensis (Coleoptera: Cerambycidae) [J]. Scientia Silvae Sinicae, 2019, 55(5): 104-113. |
| [4] | Xinlei Li,Jiatong Wang,Zhenyuan Sun,Hengfu Yin,Zhengqi Fan,Jiyuan Li. Anthocyanin Components and Their Relationship with Flower Colors in Petals of Camellia japonica 'Chidan' and Its Bud Mutation Cultivars [J]. Scientia Silvae Sinicae, 2019, 55(10): 19-26. |
| [5] | Zhang Enliang, Ma Lingling, Yang Rutong, Li Linfang, Wang Qing, Li Ya, Wang Peng. Transcriptome Profiling of IBA-Induced Adventitious Root Formation in Softwood Cuttings of Catalpa bungei ‘Yu-1’ [J]. Scientia Silvae Sinicae, 2018, 54(5): 48-61. |
| [6] | Zhang Qin, Xu Zongda, Zhao Kai, Li Xiaowei, Zhang Luosha, Zhang Qixiang. Isolation and Biological Function Analysis of Anthocyanin Regulatory Gene PmMYB1 from Prunus mume [J]. Scientia Silvae Sinicae, 2018, 54(10): 64-72. |
| [7] | Cao Yabing, Zhai Xiaoqiao, Deng Minjie, Zhao Zhenli, Fan Guoqiang. Relationship between Metabolites Variation and Paulownia Witches' Broom [J]. Scientia Silvae Sinicae, 2017, 53(6): 85-93. |
| [8] | Mao Weibing, Chen Faju, Wang Changlan, Liang Hongwei. Transcriptome Sequencing and Analysis of Male Sterile Flower Buds in Catalpa bungei [J]. Scientia Silvae Sinicae, 2017, 53(6): 141-150. |
| [9] | Yu Jian, Zhao Aichun, Liu Changying, Liang Yanmei, Zhu Panpan, Cai Yuxiang, Wang Xiling, Li Zhengang, Yu Maode. Effects of Exogenous Ethylene and 1-MCP Treatments on the Expression of Genes Involved in Ethylene and Anthocyanin in Mulberry Fruit [J]. Scientia Silvae Sinicae, 2017, 53(2): 138-148. |
| [10] | Shi Xiaodong, Zhu Xuehui, Sheng Yuzhen, Zhuang Guoqing, Chen Fang. Development of SSR Markers Based on Transcriptome Sequence of Phoebe zhennan [J]. Scientia Silvae Sinicae, 2016, 52(11): 71-78. |
| [11] | Zhang Zhen, Zhang Hanguo, Mo Chi, Zhang Lei. Transcriptome Sequencing Analysis and Development of EST-SSR Markers for Pinus koraiensis [J]. Scientia Silvae Sinicae, 2015, 51(8): 114-120. |
| [12] | Chen Hao, Tan Xiaofeng. Identification of α-Linolenic Acid Metabolism Pathway Based on Transcriptome Data of Vernicia fordii Kernels during Tung Oil Synthesis Stage [J]. Scientia Silvae Sinicae, 2015, 51(3): 41-48. |
| [13] | Wen Yafeng, Han Wenjun, Zhou Hong, Xu Gangbiao. SSR Mining and Development of EST-SSR Markers for Cunninghamia lanceolata Based on Transcriptome Sequences [J]. Scientia Silvae Sinicae, 2015, 51(11): 40-49. |
| [14] | Cui Chaoyu, Wang Yuanxiu, Jiang Junxi, Ouyang Hui, Qin Shuanglin, Huang Ting. Identification of the Pathogen Causing Brown Spot Disease of Acer rubrum ‘October Glory’ [J]. Scientia Silvae Sinicae, 2015, 51(10): 142-147. |
| [15] | Hu Yulin, Yao Xiaohua, Ren Huadong, Wang Kailiang, Lin Ping. Sequencing of Transcriptome Relevant to Flowering and Analysis of Floral-Related Genes Expression in Camellia oleifera [J]. Scientia Silvae Sinicae, 2014, 50(9): 36-43. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||