

Scientia Silvae Sinicae ›› 2022, Vol. 58 ›› Issue (7): 93-102.doi: 10.11707/j.1001-7488.20220710
• Research papers • Previous Articles Next Articles
Chuanmei Xu,Zihan Wang,Feng Xie,Xuyin Jin,Xinyan Wu,Rong Chen,Jianqin Huang*
Received:2021-09-18
Online:2022-07-25
Published:2022-11-03
Contact:
Jianqin Huang
CLC Number:
Chuanmei Xu,Zihan Wang,Feng Xie,Xuyin Jin,Xinyan Wu,Rong Chen,Jianqin Huang. Comparison of 45S rDNA and 5S rDNA Distribution Characteristics of 12 Bamboo Species[J]. Scientia Silvae Sinicae, 2022, 58(7): 93-102.
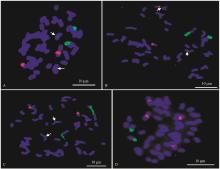
Fig.1
The distribution of 45S rDNA (green) and 5S rDNA (red) on the chromosome of four different bamboo species A to D is Phyllostachys edulis, Phyllostachys edulis 'Tao kiang', Phyllostachys edulis 'Heterocycla' and Phyllostachys violascens, respectively, white arrows show the weak signals of 5S rDNA."
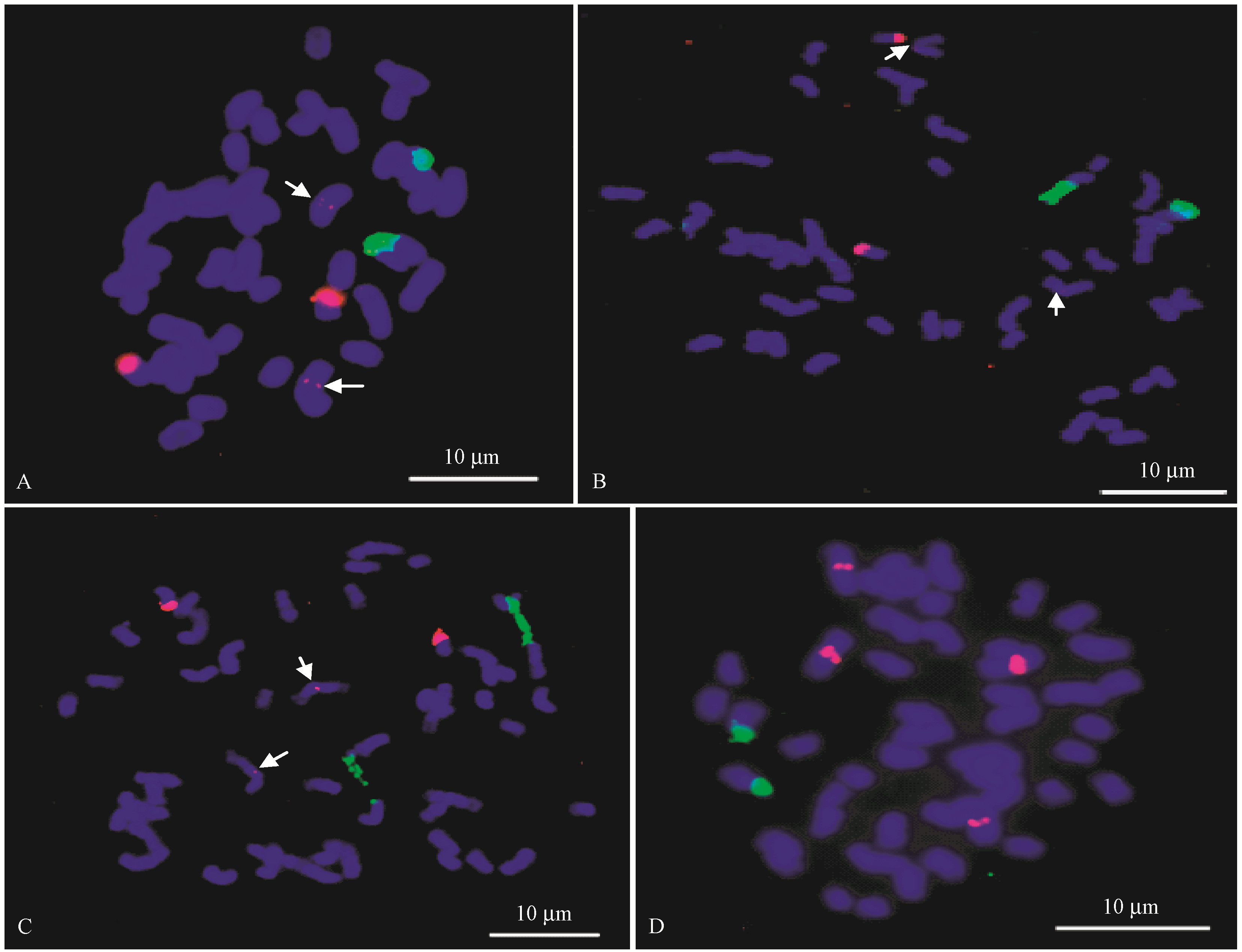
Table 1
The distribution of 45S rDNA and 5S rDNA of 12 bamboo species"
| 竹种名称 Species | 45S rDNA分布特点 Distribution of 45S rDNA | 5S rDNA分布特点 Distribution of 5S rDNA |
| 毛竹 P. edulis | 1对末端信号 A pair of terminal signals | 2对信号,1对强信号位于染色体末端,1对微弱信号位于另1对染色体近中间区域 Two pairs of signals, the pair of strong signals are located at the end of the chromosome, another pair of weak signals are located near the middle region of the chromosome |
| 花毛竹 P. edulis ‘Tao kiang’ | 1对末端信号 A pair of terminal signals | 2对信号,1对强信号位于染色体末端,1对微弱信号位于另1对染色体近中间区域 Two pairs of signals, the pair of strong signals are located at the end of the chromosome, another pair of weak signals are located near the middle region of the chromosome |
| 龟甲竹 P. edulis ‘Heterocycla’ | 1对近末端信号 A pair of subterminal signals | 2对信号,1对强信号位于染色体末端,1对微弱信号位于另1对染色体近中间区域 Two pairs of signals, the pair of strong signals are located at the end of the chromosome, another pair of weak signals are located near the middle region of the chromosome. |
| 早竹 P. violascens | 1对末端信号 A pair of terminal signal | 2对较强的信号,均位于染色体近中间位置 Two pairs of strong signals are located near the middle region of the chromosome |
| 白哺鸡竹 P. dulcis | 1对末端信号, 其中一个位点发生了一定断裂 A pair of terminal signals, one of them is broken | 2对较强的信号,均位于染色体近中间位置 Two pairs of strong signals are located near the middle region of the chromosome. |
| 黄秆乌哺鸡竹P. vivax f. aureocaulis | 1对末端信号,存在一定的断裂 A pair of broken terminal signals | 2对信号,均位于染色体近中间位置 Two pairs of signals are located near the middle region of the chromosome |
| 斑竹 P. reticulata ‘Lacrima-deae’ | 1对末端信号,存在一定的断裂 A pair of broken terminal signals | 2对信号,均位于染色体近中间位置 Two pairs of signals are located near the middle region of the chromosome |
| 人面竹 P. aurea | 1对末端信号,其中一个位点发生了一定断裂 A pair of terminal signals, one of them is broken | 2对信号,1对信号强度相似,另1对信号表现为1强1弱 Two pairs of signals, one pair are similar, another pair are different in intensity |
| 金镶玉竹 P. aureosulcata ‘Spectabilis’ | 1对末端信号,其中一个位点发生了一定断裂 A pair of terminal signals, one of them is broken | 3个信号,2个位于染色体近中间位置,表现1强1弱,1个位于染色体末端,信号较强 Three different signals, two of them are different in intensity and located near the middle region of the chromosome, the last one is strong and located at the end of the chromosome |
| 紫竹 P. nigra | 1强1弱2个信号,强信号易断裂 Two different strength signals, the strong one is easy to break. | 3个信号,1对信号很强,位于染色体末端,另1个信号较弱,位于染色体近中间位置 Three signals, a strong pair are located at the end of the chromosome, another weak signal is located near the middle region of the chromosome |
| 茶秆竹 P. amabilis | 1对近末端信号,其中一个位点易断裂成2段 A pair of terminal signals, one of them easily breaks into 2 segments | 1对较弱的信号,位于染色体短臂近着丝粒部位 A pair of weak signals are located near the centromere of the short arm |
| 大明竹 P. gramineus | 2个不同的末端信号 Two different terminal signals | 3个信号,2个位于染色体短臂中间,另1个位于染色体近中间区域 Three signals, two are located at the middle of the short arm, the third located near the middle region of the chromosome |
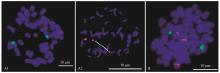
Fig.4
The distribution of 45S rDNA (green) and 5S rDNA (red) on the chromosome of two different bamboo species A1 and B is the distribution of 45S rDNA (green) and 5S rDNA(red)on the chromosome of Pseudoasa amabilis and Pleioblastus gramineus, respectively. A2 is the distribution of 45S rDNA (red) on the Pseudoasa amabilis."
| 陈瑞阳, 李秀兰, 宋文芹, 等. 中国主要经济植物基因组染色体图谱Ⅳ. 北京: 科学出版社, 2003: 1- 628. | |
| Chen R Y , Li X L , Song W Q , et al. Chromosome atlas of major economic plants genome in china (Version IV): chromosome atlas of various bamboo species. Beijing: Science Press, Science Press, 2003: 1- 628. | |
| 贾芳信, 周明兵, 陈荣, 等. 4种竹子的核型及其基因组大小. 林业科学, 2016, 52 (9): 57- 66. | |
| Jia F X , Zhou M B , Chen R , et al. Karyotype and genome size in four bamboo species. Scientia Silvae Sinicae, 2016, 52 (9): 57- 66. | |
| 李秀兰, 林汝顺, 冯学琳, 等. 中国部分丛生竹类染色体数目报道. 植物分类学报, 2001, 39 (5): 433- 422. | |
| Li X L , Lin R S , Feng R S , et al. Chromosome number of some caespitose bamboos native in or introduced to China. Acta Phytotaxonomica Sinica, 2000, 39 (5): 433- 422. | |
| 李秀兰, 刘松, 宋文芹, 等. 40种散生竹的染色体数目. 植物分类学报, 1999, 37 (6): 541- 544. | |
| Li X L , Liu S , Song W Q , et al. Chromosome number of forty species of scattered bamboos. Acta Phytotaxonomica Sinica, 1999, 37 (6): 541- 544. | |
| 马乃训, 赖广辉, 张培新, 等. 中国刚竹属. 杭州: 浙江科学技术出版社, 2014: 28- 154. | |
| Ma N X , Lai G H , Zhang P X , et al. The genus Phyllostachys in China. Hangzhou: Zhejiang Science and Technology Press, 2014: 28- 154. | |
| 徐川梅, 卢江杰, 汤定钦. 45S rDNA在7种竹子植物染色体上的定位. 林业科学, 2009a, 45 (12): 42- 45. | |
| Xu C M , Lu J J , Tang D Q . Physical mapping of the 45S rDNA on metaphase chromosomes in seven bamboo species by in situ hybridization. Scientia Silvae Sinicae, 2009a, 45 (12): 42- 45. | |
| 徐川梅, 郑华威, 王骢, 等. 龟甲竹染色体C-分带、荧光原位杂交及其核型分析. 林业科学研究, 2009b, 22 (5): 691- 695. | |
| Xu C M , Zhen H W , Wang C , et al. Giemsa C-banding, FISH and karyotype analysis of Phyllostachys pubescens var.heterocycla chromosome. Forest Research, 2009b, 22 (5): 691- 695. | |
| 张颖, 赵欣, 张圣虎, 等. 竹类植物修复重金属污染土壤的研究进展. 生态与农村环境学报, 2021, 37 (1): 30- 38. | |
| Zhang Y , Zhao X , Zhang S H , et al. Research progress of bamboo species in phytoremediation of heavy metal contaminated soil. Journal of Ecology and Rural Environment, 2021, 37 (1): 30- 38. | |
|
Badaeva E D , Dedkova O S , Gay G , et al. Chromosomal rearrangements in wheat: their types and distribution. Genome, 2007, 50 (10): 907- 926.
doi: 10.1139/G07-072 |
|
|
Garcia S , Kovarík A , Leitch A R , et al. Cytogenetic features of rRNA genes across land plants: analysis of the plant rDNA database. Plant Journal, 2017, 89 (5): 1020- 1030.
doi: 10.1111/tpj.13442 |
|
|
Guo Z H , Ma P F , Yang G Q , et al. Genome sequences provide insights into the reticulate origin and unique traits of woody bamboos. Molecular Plant, 2019, 12 (10): 1353- 1365.
doi: 10.1016/j.molp.2019.05.009 |
|
|
Hasterok R , Wolny E , Hosiawa M , et al. Comparative analysis of rDNA distribution in chromosomes of various species of Brassicaceae. Annals of Botany, 2006, 97 (2): 205- 216.
doi: 10.1093/aob/mcj031 |
|
|
Heslop-Harrison J S . Comparative genome organization in plants: from sequence and markers to chromatin and chromosomes. Plant Cell, 2000, 12 (5): 617- 636.
doi: 10.1105/tpc.12.5.617 |
|
| Huang J , Ma L , Yang F , et al. 45S rDNA regions are chromosome fragile sites expressed as gaps in vitro on metaphase chromosomes of root-tip meristematic cells in Lolium spp. PLoS One, 2008, 3 (5): e2167. | |
| Jiang J , Gill B S , Wang G L , et al. Metaphase and interphase fluorescence in situ hybridization mapping of the rice genome with bacterial artificial chromosomes. Proceedings of the National Academy of Sciences, 1995, 92 (10): 4487- 4491. | |
| Katsiotis A , Loukas M , Heslop-Harrison J S . Repetitive DNA, genome and species relationships in Avena and Arrhenatherum (Poaceae). Annals of Botany, 2000, 86 (6): 1135- 1142. | |
| Luo X M , Liu J C . Fluorescence in situ hybridization (FISH) analysis of the locations of the oligonucleotides 5S rDNA, (AGGGTTT)3, and (TTG) 6 in three genera of Oleaceae and their phylogenetic framework. Genes, 2019, 10 (5): 375. | |
| Mondin M , Aguiar-Perecin , M L R . Heterochromatin patterns and ribosomal DNA loci distribution in diploid and polyploid Crotalaria species (Leguminosae, Papilionoideae) and inferences on karyotype evolution. Genome, 2011, 54 (9): 718- 726. | |
| Monkheang P , Chaveerach A , Sudmoon R , et al. Karyotypic features including organizations of the 5S, 45S rDNA loci and telomeres of Scadoxus multiflorus (Amaryllidaceae). Comparative Cytogenetics, 2016, 10 (4): 637- 646. | |
| Osuji J O , Crouch J , Harrison G , et al. Molecular cytogenetics of Musa species, cultivars and hybrids: location of 18S-5.8S-25S and 5S rDNA and telomere-like sequences. Annals of Botany, 1998, 82 (2): 243- 248. | |
| Pedrosa A , Sandal N , Stougaard J , et al. Chromosomal map of the model legume Lotus japonicas. Genetics, 2002, 161 (4): 1661- 1672. | |
| Pedrosa-Harand A , de Almeida C C S , Mosiolek M , et al. Extensive ribosomal DNA amplification during Andean common bean (Phaseolus vulgaris L.) evolution. Theoretical and Applied Genetics, 2006, 112 (5): 924- 33. | |
| Peng Z , Lu Y , Li L , et al. The draft genome of the fast-growing non-timber forest species moso bamboo (Phyllostachys heterocycla). Nature Genetics, 2013, 45 (4): 456- 461. | |
| Raskina O , Barber J C , Nevo E , et al. Repetitive DNA and chromosomal rearrangements: speciation related events in plant genomes. Cytogenetic and Genome Research, 2008, 120 (3/4): 351- 357. | |
| Tsang E , Carr A M . Replication fork arrest, recombination and the maintenance of ribosomal DNA stability. DNA Repair, 2008, 7 (10): 1613- 1623. | |
| Vaio M , Speranza P , Valls J F , et al. Localization of the 5S and 45S rDNA sites and cpDNA sequence analysis in species of the Quadrifaria group of Paspalum (Poaceae, Paniceae). Annals of Botany, 2005, 96 (2): 191- 200. | |
| Vasconcelos E V , Vasconcelos S , Ribeiro T , et al. Karyotype heterogeneity in Philodendron s.l. (Araceae) revealed by chromosome mapping of rDNA loci. PLoS One, 2018, 13 (11): 1- 15. | |
| Zhang L N , Ma P F , Zhan Y X , et al. Using nuclear loci and allelic variation to disentangle the phylogeny of Phyllostachys (Poaceae, Bambusoideae). Molecular Phylogenetics and Evolution, 2019, 137, 222- 235. | |
| Zhang Z T , Yang S Q , Li Z A , et al. Comparative chromosomal localization of 45S and 5S rDNAs and implications for genome evolution in Cucumis. Genome, 2016, 59 (7): 449- 457. | |
| Zhao H , Gao Z , Wang L , et al. Chromosome-level reference genome and alternative splicing atlas of moso bamboo (Phyllostachys edulis). Gigascience, 2018, 7 (10): 1- 12. | |
| Zhou M B , Xu C M , Shen L F , et al. Evolution of genome sizes in Chinese Bambusoideae (Poaceae) in relation to karyotype. Trees, 2017, 31, 41- 48. |
| [1] | Yanmin Li,Deyi Yuan,Tianwen Ye,Ya Chen,Chunxia Han,Shixin Xiao. Karyotype Analysis of 18 Excellent Individuals in F1 Generation of Interspecific Hybridization of Oil-Tea (Camellia oleifera) [J]. Scientia Silvae Sinicae, 2022, 58(4): 165-174. |
| [2] | Lüji Wang,Xiaoxiao Zhang,Jun Wang. Chromosome Behaviors Tracking during Meiosis of Microsporocytes of Allotriploid Populus alba × P. berolinensis 'Yinzhong' Based on 45S rDNA-FISH [J]. Scientia Silvae Sinicae, 2022, 58(3): 69-78. |
| [3] | Tianwen Ye,Deyi Yuan,Yanmin Li,Shixin Xiao,Shoufu Gong,Jian Zhang,Sufang Li,Jian Luo. Ploidy Identification of Camellia hainanica [J]. Scientia Silvae Sinicae, 2021, 57(7): 61-69. |
| [4] | Ruihong Guo,Shuai Qiu,Guangxin Liu,Kai Gao,Jianfen Wei,Mengli Xi. Optimization of Chromosome Preparation and rDNA Physical Localization of Hydrangea macrophylla [J]. Scientia Silvae Sinicae, 2021, 57(11): 59-67. |
| [5] | Xu Zilong, Chen Yicun, Gao Ming, Wu Liwen, Zhao Yunxiao, Wang Yangdong. Research Progress in Sex Differentiation in Angiosperms [J]. Scientia Silvae Sinicae, 2019, 55(8): 157-169. |
| [6] | Wang Jirui, Zheng Chenyi, Du Yuzhou, Wang Hao, Zhou Guoxin, Xu Zhihong. Three Species of Whiteflies(Hemiptera: Aleyrodidae) Infesting Bamboo in China [J]. Scientia Silvae Sinicae, 2018, 54(7): 173-180. |
| [7] | Tian Mengdi, Li Yanjie, Zhang Pingdong, Wang Jian, Hao Jingyi. Pollen Chromosome Doubling Induced by High Temperature Exposure to Produce Hybrid Triploids in Populus canescens [J]. Scientia Silvae Sinicae, 2018, 54(3): 39-47. |
| [8] | Xu Chuanmei, Huang Jianqin, Wang Zhengjia, Xia Guohua, Zhang Qixiang, Huang Youjun, Zhang Shougong. Meiosis of Pollen Mother Cell and Karyotype of Carya cathayensis [J]. Scientia Silvae Sinicae, 2017, 53(11): 77-84. |
| [9] | Jia Fangxin, Zhou Mingbing, Chen Rong, Yang Haiyun, Gao Peijun, Xu Chuanmei. Karyotype and Genome Size in Four Bamboo Species [J]. Scientia Silvae Sinicae, 2016, 52(9): 57-66. |
| [10] | Li Tiezhu;Tian Dalun;Wuyun Tana;Tan Xiaofeng;Xiang Wenhua;Yan Wende;. Identification and Variation of Tetraploid Camellia oleifera [J]. Scientia Silvae Sinicae, 2009, 12(3): 150-154. |
| [11] | Xu Chuanmei;Lu Jiangjie;Tang Dingqin. Physical Mapping of the 45S rDNA on Metaphase Chromosomes in Seven Bamboo Species by in situ Hybridization [J]. Scientia Silvae Sinicae, 2009, 12(12): 42-45. |
| [12] | Wang Xiaorong;Tang Haoru;Fu Huaqing;Zhong Bifeng;Xia Wufeng;Tang Haidong;Deng Qunxian. Karyotypes of 15 Introduced Bramble Cultivars (Rubus) [J]. Scientia Silvae Sinicae, 2008, 44(12): 147-150. |
| [13] | Xu Jin;Shi Jisen;Yang Liwei;Wang Guifeng. Cell Biological Characteristics and Abnormal Behavior during the Meiosis of Pollen Mother Cells in Cunninghamia lanceolata [J]. Scientia Silvae Sinicae, 2007, 43(11): 32-36. |
| [14] | Yin Aihua;Jin Hui;Han Zhengmin;Han Sufen. Study on the Relationship between Chromosome Numbers and Nodulation of 18 Species of Leguminous Trees [J]. Scientia Silvae Sinicae, 2006, 42(1): 26-28. |
| [15] | Yumei Wang,Maoxue Li,Ruilin You. OBSERVATION ON CHROMOSOMES OF GINKGO BILOBA BY USING MATERIALS RATHER THAN ROOT-TIP CELLS [J]. Scientia Silvae Sinicae, 1999, 35(5): 2-4. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||