

Scientia Silvae Sinicae ›› 2021, Vol. 57 ›› Issue (3): 170-180.doi: 10.11707/j.1001-7488.20210318
• Scientific notes • Previous Articles Next Articles
Minhao Liu,Long Li,Jing Ye,Xuanyuan Zhou,Zhouqi Li*,Ruishen Fan,Junlei Xu
Received:2020-05-27
Online:2021-03-25
Published:2021-04-07
Contact:
Zhouqi Li
CLC Number:
Minhao Liu,Long Li,Jing Ye,Xuanyuan Zhou,Zhouqi Li,Ruishen Fan,Junlei Xu. Genome-Wide Identification and Expression Analysis of the ARF Gene Family in Eucommia ulmoides[J]. Scientia Silvae Sinicae, 2021, 57(3): 170-180.
Table 1
eu-miR160s/eu-miR167s sequences and primers"
| miRNA | 序列Sequences (5′—3′) | 茎环引物Stem ring primer (5′—3′) | 正向引物Forward primer (5′—3′) |
| eu-miR160d | TGCCTGGCTCCCTGCATGCCA | CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGCTGGCATGC | TCGGCAGGTGCCTGGCTCCCT |
| eu-miR160f | TGCCTGGCTCCCTGTATGCC | CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGCGGCATACA | TCGGCAGGTGCCTGGCTCCC |
| eu-miR167a | TGAAGCTGCCAGCATGATCTC | CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGCGAGATCAT | TCGGCAGGTGAAGCTGCCAGC |
| eu-miR167d | TGAAGCTGCCAGCATGATCT | CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGCAGATCATG | TCGGCAGGTGAAGCTGCCAG |
Table 2
EuARFs gene primers"
| 基因 Gene | 正向引物Forward primer (5′—3′) | 反向引物Reverse primer (5′—3′) |
| EuARF1 | CTTCAGGTGCCAAGCCTAGTTCTC | GCCGGTCTTCTTATGCCAGTGTC |
| EuARF2.1 | CCATAGTGAACAGGTGGCAGCATC | GGTTGGAGCGTCATCTGAGCATAC |
| EuARF2.2 | GCATGGCATACTTGCGAGTG | CCGAACGAGTAATCATATTCAACAG |
| EuARF3.1 | ACGAAGGTGACATGATGCTGATGG | TCGTTGCCGCTGCCAATTCC |
| EuARF3.2 | AGCCAGCACCATCACTTCC | AGCCAGCACCATCACTTCC |
| EuARF4 | GAAGGTTCAGCGGTCTTGTG | TCCCATTTCACTATCAAGCATCTC |
| EuARF6.1 | TCGCCACAAGAGGTGCAGGAG | GCATGGTCATCGCAGCTACTATCG |
| EuARF6.2 | GATACTATCGCCGCTGGAAGTGC | TGTCGCCGTTGCCGTGATTAAG |
| EuARF8.1 | CATTGGAGATCCGTCAAGGTTGGTT | ATGGGAAAGGTTGTCAAAGGCTCAA |
| EuARF8.2 | GGGCTTGTCCATCTGAGTTTG | GCACCAGTTTGACGACCTTC |
| EuARF9 | TCCACAGCAACAGATGACACAACC | GCTGCGGAAGCTGAGAGTATAACG |
| EuARF10.1 | GCGTCCTGGGATGAGGTTTA | CTGAAGACACTGCCATGAAGAAT |
| EuARF10.2 | TTGGAACAGACGATACAGCAGACAA | GAGCCAGGCCATCTAGCAGGAT |
| EuARF16 | TCGTCAACCGCAAGAAGCTCATC | GCGTTGGCTGAAGGTAGAGGAAG |
| EuARF17 | ATGTGAAGCGTGTCAGCCCTTG | AGGTGGTGAGAACTGTGGAATCTGA |
| EuARF18 | ACCGACAGCGAGAATGACATGATG | GCCGACAGATGAAGACTTGGACAG |
| EuARF19.1 | GGTGCGGTGGGATGAAACTTCTAC | CAGGAGTCAGGGCAGGCTCTATT |
| EuARF19.2 | TTGCCTCAGGTTGGAAGTCTTGTC | GTCTGCGTGTAATGTGGCGTTATG |
Table 3
EuARF gene family information"
| 基因 Gene | 基因编号 Gene ID | 蛋白长度 Protein length (aa) | 等电点 pI | 分子量 Molecular weight/ kDa | 结构域 Domain |
| EuARF1 | GWHGAAAL013439 | 668 | 6.15 | 74.75 | DBD, MR, CTD |
| EuARF2.1 | GWHGAAAL012635 | 822 | 7.62 | 91.97 | DBD, MR, CTD |
| EuARF2.2 | GWHGAAAL023550 | 577 | 6.18 | 65.12 | DBD, MR, CTD |
| EuARF3.1 | GWHGAAAL026509 | 638 | 6.27 | 70.13 | DBD, ARF |
| EuARF3.2 | GWHGAAAL025681 | 823 | 6.69 | 90.37 | DBD, ARF |
| EuARF4 | GWHGAAAL008814 | 846 | 5.91 | 94.44 | DBD, MR, CTD |
| EuARF6.1 | GWHGAAAL000541 | 892 | 5.75 | 99.40 | DBD, MR, CTD |
| EuARF6.2 | GWHGAAAL008938 | 858 | 5.76 | 95.04 | DBD, MR, CTD |
| EuARF8.1 | GWHGAAAL000541 | 892 | 5.75 | 99.40 | DBD, MR, CTD |
| EuARF8.2 | GWHGAAAL000012 | 273 | 9.68 | 30.64 | DBD, ARF |
| EuARF9 | GWHGAAAL016769 | 653 | 5.68 | 73.19 | DBD, MR, CTD |
| EuARF10.1 | GWHGAAAL008845 | 631 | 6.19 | 69.97 | DBD, ARF |
| EuARF10.2 | GWHGAAAL008846 | 633 | 5.36 | 70.70 | DBD, ARF |
| EuARF16 | GWHGAAAL011358 | 682 | 6.96 | 74.95 | DBD, ARF |
| EuARF17 | GWHGAAAL001628 | 526 | 5.63 | 57.54 | DBD, ARF |
| EuARF18 | GWHGAAAL025932 | 623 | 6.04 | 70.23 | DBD, MR, CTD |
| EuARF19.1 | GWHGAAAL003011 | 1 062 | 6.05 | 117.84 | DBD, MR, CTD |
| EuARF19.2 | GWHGAAAL003601 | 1 013 | 6.12 | 111.39 | DBD, MR, CTD |
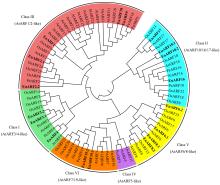
Fig.1
Phylogenetic tree of the ARF gene family Phylogenetic relationships of Arabidopsis thaliana (AtARF, 22), Oryza sativa (OsARF, 25), Populus × euramericana (PeARF, 14) and Eucommia ulmoides (EuARF, 18) ARF proteins constructed by MEGA5.1 according to Neighbor-Joining and the bootstrap valued at 1 000."
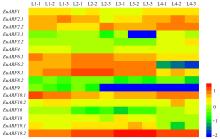
Fig.5
Expression profiles of EuARF genes in leaves at different growth stages Color scale represents FPKM normalized log10 function transformed counts. L1 is the leaf bud, L2 is the 3 cm long growing leaf, L3 is the fully expanded young leaf, and L4 is the leaf growing 30 days after fully expanded. The number after"-" means three repeated tests performed on each sample."
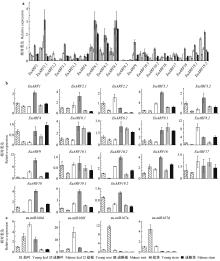
Fig.6
Expression of EuARFs genes and eu-miR160s/eu-miR167s in organs at different developmental stages The EuUBC gene is used as an internal control. a. The expression of EuARF6 in the young leaf is used as a control; b. The expression of each EuARF in the young leaf is used as a control; c. The expression of each miRNA in the young leaf is used as control. Bars represent standard deviations for three biological replicates."
|
董晨, 魏永赞, 王弋, 等. 基于转录组的荔枝ARF基因家族的鉴定及表达分析. 热带作物学报, 2017, 38 (8): 1485- 1491.
doi: 10.3969/j.issn.1000-2561.2017.08.018 |
|
|
Dong C , Wei Y Z , Wang Y , et al. Transcriptome-wide identification and expression profiling of the auxin response factor(ARF) gene family in Litchi chinensis Sonn. Chinese Journal of Tropical Crops, 2017, 38 (8): 1485- 1491.
doi: 10.3969/j.issn.1000-2561.2017.08.018 |
|
| 张京京, 杜红岩, 李钦, 等. 杜仲药理与毒理研究进展. 河南大学学报: 医学版, 2014, 33 (3): 217- 222. | |
| Zhang J J , Du H Y , Li Q , et al. Study advancement about pharmacological and toxicological of Eucommia ulmoides Oliv. Journal of Henan University: Medical Science, 2014, 33 (3): 217- 222. | |
|
Hagen G , Guilfoyle T . Auxin-responsive gene expression: genes, promoters and regulatory factors. Plant Molecular Biology, 2002, 49, 373- 385.
doi: 10.1023/A:1015207114117 |
|
|
Hong Y , Marçal S , Isabelle M , et al. Genome-wide characterization and expression profiling of the auxin response factor(ARF) gene family in Eucalyptus grandis. PLoS ONE, 2014, 9, e108906.
doi: 10.1371/journal.pone.0108906 |
|
|
Huseyin T . Genome-wide analysis of the auxin response factors(ARF) gene family in barley(Hordeum vulgare L.). Journal of Plant Biochemistry and Biotechnology, 2018, 28, 14- 24.
doi: 10.1007/s13562-018-0458-6 |
|
|
Kalluri U C , DiFazio S P , Brunner A M , et al. Genome-wide analysis of Aux/IAA and ARF gene families in Populus trichocarpa. BMC Plant Biology, 2007, 7 (1): 59- 69.
doi: 10.1186/1471-2229-7-59 |
|
| Li S B , Ouyang W Z , Hou X J , et al. Genome-wide identification, isolation and expression analysis of auxin response factor(ARF) gene family in sweet orange (Citrus sinensis). Frontiers in Plant Science, 2015, 6, 119. | |
| Lin Y , Lai Z , Tian Q , et al. Endogenous target mimics down-regulate miR160 mediation of ARF10, -16 and -17 cleavage during somatic embryogenesis in Dimocarpus longan Lour. Front Plant Sci, 2015, 6, 956. | |
| Liu Z H , Yu Y C . Auxin response factors and plant growth and development. Hereditas, 2011a, 33 (12): 1335- 1346. | |
| Liu R E , Hu C G , Sun Y Q . Advances in plant auxin response factors. Plant Physiology Journal, 2011b, 47 (7): 669- 679. | |
|
Luo X C , Sun M H , Xu R R , et al. Genomewide identification and expression analysis of the ARF gene family in apple. Journal of Genetics, 2014, 93 (3): 785- 797.
doi: 10.1007/s12041-014-0462-0 |
|
|
Okushima Y , Mitina I , Quach H L , et al. Auxin response factor 2(ARF2): a pleiotropic developmental regulator. The Plant Journal, 2005, 43 (1): 29- 46.
doi: 10.1111/j.1365-313X.2005.02426.x |
|
|
Okushima Y , Fukaki H , Onoda M , et al. ARF7 and ARF19 regulate lateral root formation via direct activation of LBD/ASL genes in Arabidopsis. Plant Cell, 2007, 19 (1): 118- 130.
doi: 10.1105/tpc.106.047761 |
|
|
Tang Y H , Bao X X , Liu K , et al. Genome-wide identification and expression profiling of the auxin response factor(ARF) gene family in physic nut. PLoS ONE, 2018, 13, e0201024.
doi: 10.1371/journal.pone.0201024 |
|
|
Tiwari S B . The roles of auxin response factor domains in auxin-responsive transcription. The Plant Cell Online, 2003, 15 (2): 533- 543.
doi: 10.1105/tpc.008417 |
|
|
Wan S , Li W , Zhu Y , et al. Genome-wide identification, characterization and expression analysis of the auxin response factor gene family in Vitis vinifera. Plant Cell Reports, 2014, 33 (8): 1365- 1375.
doi: 10.1007/s00299-014-1622-7 |
|
| Wang D , Pei K , Fu Y , et al. Genome-wide analysis of the auxin response factors (ARF) gene family in rice (Oryza sativa). Gene, 2007, 394 (1/2): 13- 24. | |
|
Wang J W , Wang L J , Mao Y B , et al. Control of root cap formation by microRNA-targeted auxin response factors in Arabidopsis. Plant Cell, 2005, 17, 2204- 2216.
doi: 10.1105/tpc.105.033076 |
|
|
Wang Y , Li K , Chen L , et al. MicroRNA167-directed regulation of the auxin response factors, GmARF8a and GmARF8b, is required for soybean(Glycine max L.) nodulation and lateral root development. Plant Physiol, 2015, 168 (3): 984- 999.
doi: 10.1104/pp.15.00265 |
|
| Williams L , Carles C C , Osmont K S , et al. A database analysis method identifies an endogenous transacting short-interfering RNA that targets the Arabidopsis ARF2, ARF3, and ARF4 genes. Proc Natl Acad Sci USA, 2005, 102 (77): 9703- 9708. | |
|
Wu J , Wang F , Cheng L , et al. Identification, isolation and expression analysis of auxin response factor(ARF) genes in Solanum lycopersicum. Plant Cell Reports, 2011, 30 (11): 2059- 2073.
doi: 10.1007/s00299-011-1113-z |
|
|
Wu M F , Tian Q , Reed J W . Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development, 2006, 133 (21): 4211- 4218.
doi: 10.1242/dev.02602 |
|
|
Wuyun T N , Wang L , Liu H , et al. The hardy rubber tree genome provides insights into the evolution of polyisoprene biosynthesis. Molecular Plant, 2018, 11 (3): 429.
doi: 10.1016/j.molp.2017.11.014 |
|
|
Xing H , Pudake R N , Guo G , et al. Genome-wide identification and expression profiling of auxin response factor (ARF) gene family in maize. BMC Genomics, 2011, 12, 178.
doi: 10.1186/1471-2164-12-178 |
|
|
Yang C , Xu M , Xuan L , et al. Identification and expression analysis of twenty ARF genes in Populus. Gene, 2014, 544 (2): 134- 144.
doi: 10.1016/j.gene.2014.04.067 |
|
|
Ye J , Jin C F , Li L , et al. Selection of suitable reference genes for qRT-PCR normalisation under different experimental conditions in Eucommia ulmoides Oliv. Scientific Reports, 2018, 8, 15043.
doi: 10.1038/s41598-018-33342-w |
|
| Zhang S , Wang S , Xu Y , et al. The auxin response factor, OsARF19, controls rice leaf angles through positively regulating OsGH3-5 and OsBRI1. Plant, Cell & Environment, 2015, 38 (4): 638- 654. |
| [1] | Pu Zhang,Wenjing Shao,Kejiu Du,Shuang Zhang. Cloning and Functional Analysis of Cysteine Proteinase Inhibitor Gene EuCPI from Eucommia ulmoides [J]. Scientia Silvae Sinicae, 2021, 57(3): 29-38. |
| [2] | Minhao Liu,Junlei Xu,Jing Ye,Zhouqi Li,Ruishen Fan,Long Li. Agrobacterium tumefaciens-Mediated Transformation of Leaf Callus in Eucommia ulmoides [J]. Scientia Silvae Sinicae, 2020, 56(2): 79-88. |
| [3] | Zeng Zhaoyan, Li Xiangzhou, Zhang Sheng, Huang Dan. Adsorption and Sustained Release of Eucommia ulmoides Extract on Nano-Bamboo Charcoal [J]. Scientia Silvae Sinicae, 2018, 54(6): 125-131. |
| [4] | Li Hongguo, Xu Jihuang, Du Hongyan, Wuyun Tana, Liu Panfeng, Du Qingxin. Preliminary Construction of Core Collection of Eucommia ulmoides Based on Allele Number Maximization Strategy [J]. Scientia Silvae Sinicae, 2018, 54(2): 42-51. |
| [5] | Cao Ruizhi, Zhang Xinyu, Yang Dawei, Xia Guangdong, Dong Juan. Effects of Different Barking Treatments on Secondary Metabolites of Eucommia ulmoides and Its Ability of Repairing Injury [J]. Scientia Silvae Sinicae, 2017, 53(6): 151-158. |
| [6] | Du Qingxin, Wei Yanxiu, Liu Panfeng, Du Hongyan. Diversity of the Content of Main Active Components in Eucommia ulmoides Male Flowers [J]. Scientia Silvae Sinicae, 2017, 53(2): 35-43. |
| [7] | Sun Zhiqaing, Zhao Yang, Ma Zhigang, Du Hongyan, Zhu Jingle, Wang Hao, Chen Junhua, Fu Jianmin. Risk Grading for Damage of the Defoliator Orthosia songi (Lepidoptera: Noctuidae) [J]. Scientia Silvae Sinicae, 2016, 52(7): 78-86. |
| [8] | Wang Lu, Wuyun Tana, Du Lanying, Liu Panfeng, Du Hongyan. An Improved Variety for Samara and Medicinal Use: Eucommia ulmoides ‘Huazhong 10’ [J]. Scientia Silvae Sinicae, 2016, 52(11): 171-171. |
| [9] | Lin Kaiqin, Zhao Degang, Li Yan, He Xuanze, Wang Shaomin. Development of Gender-Related EST-SSR Markers in Eucommia ulmoides [J]. Scientia Silvae Sinicae, 2016, 52(10): 146-152. |
| [10] | Kangjian Zhang,Yaqin Wang,Xihan Ma,Lan Wang,Tan Zhang. AN ECOLOGICAL STUDY ON SECONDARY METABOLITES OF THE LEAVES OF EUCOMMIA ULMOIDES [J]. Scientia Silvae Sinicae, 1999, 35(6): 28-34. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||