

Scientia Silvae Sinicae ›› 2022, Vol. 58 ›› Issue (3): 59-68.doi: 10.11707/j.1001-7488.20220307
Previous Articles Next Articles
Hebi Zhuang,Zicheng Yu,Erpei Lin,Huahong Huang*
Received:2021-04-06
Online:2022-03-25
Published:2022-06-02
Contact:
Huahong Huang
CLC Number:
Hebi Zhuang,Zicheng Yu,Erpei Lin,Huahong Huang. Cloning, Expression and Interaction Protein Screening of BlBLH1 Gene in Betula luminifera[J]. Scientia Silvae Sinicae, 2022, 58(3): 59-68.
Table 1
PCR primers used in this study"
| 引物名称 Primer name | 引物序列(5′-3′) Primer sequence(5′-3′) | 用途 Application |
| BlBLH1-FF | GCATCAGAATGGGATAAAGTC | 基因克隆 Gene clone |
| BlBLH1-FR | CCATTTCCTTTTCGATCTTG | |
| BlBLH1-F | ATGAATCCCAACTACGTGCCGTA | 构建酵母杂交载体 Construction of yeast two hybrid plasmids |
| BlBLH1-R | GGCCACGAAATCTGGCAACAATTG | |
| BlKNOX4-F | ATGGAAGGAGGAGTTGCTGGTAG | |
| BlKNOX4-R | AAGGAGCATAGGGGTGCAATCCA | |
| BlKNOX9-F | ATGCATTTGTATATCCTCATCCC | |
| BlKNOX9-R | GTCATTCCTGAAAAGCGGTCCAT | |
| BlSTM-F | ATGGAGAGTGGCTCTAATG | |
| BlSTM-R | GAGCAGTGTGGGCGAAATAT | |
| BlEF1α-QF | GAATAAGATGGATGAGCCTACCGT | 荧光定量PCR Quantitative real-time PCR |
| BlEF1α-QR | CAGACCAGATATTGGCAGGAACT | |
| BlBLH1-QF | ACCCCACAACCATCTCATTC | |
| BlBLH1-QR | AGTTCTTCAATCAGGCCACG | |
| BlKNOX4-QF | CATTGGAAGCCTTCAGAAGATATG | |
| BlKNOX4-QR | TACAGTGACCAGGAAAATGTCATC | |
| BlKNOX9-QF | GCTCAAGCTTCGATGGATGAACTG | |
| BlKNOX9-QR | TGCAGAATTTCAGTCATTCCTGAG | |
| BlBLH1-inF | AGATCTCGAGCTCAAGCTTCGATGAATCCCAACTACGTGCCG | 构建BiFC载体 Construction of BiFC plasmids |
| BlBLH1-inR | GGTACCGTCGACTGCAGAATTTCAGGCCACGAAATCTGGCAAC | |
| BlKNOX4-inF | AGATCTCGAGCTCAAGCTTCGATGGAAGGAGGAGTTGCTGGTAG | |
| BlKNOX4-inR | GGTACCGTCGACTGCAGAATTTCAAAGGAGCATAGGGGTGCAAT | |
| BlKNOX9-inF | AGATCTCGAGCTCAAGCTTCGATGGATGAACTGTATGGCATC | |
| BlKNOX9-inR | GGTACCGTCGACTGCAGAATTTCAGTCATTCCTGAAAAGCG | |
| BlSTM-inF | AGATCTCGAGCTCAAGCTTCGATGGAGAGTGGCTCTAATG | |
| BlSTM-inR | GGTACCGTCGACTGCAGAATTTCAGAGCAGTGTGGGCGAAAT |
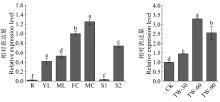

Fig.3
Expression analysis of BlBLH1 in different tissues, organs and tension wood formation Different small letters above columns indicate significant difference at 0. 05 levels. R: Root; YL: Young leaf; ML: Mature leaf; FF: Female inflorescence; MF: Male inflorescence; S1: Non-lignified stem; S2: Lignified stem; TW: Tension wood."
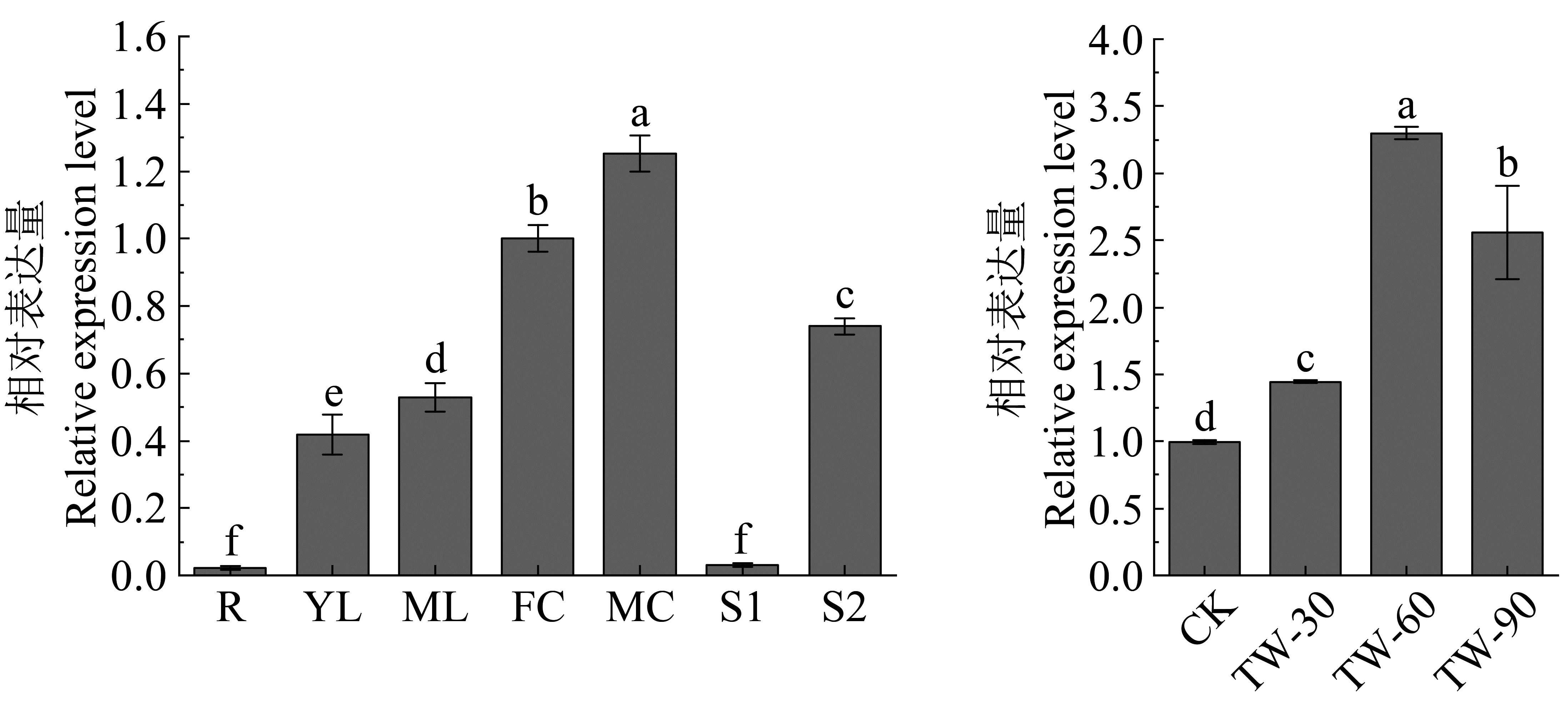
Table 2
The BLAST results of BlBLH1 yeast library hybridization positive clones"
| 编号 No. | 基因编号 Gene_ID | 同源基因序列号 Homolog No. | 注释 Description |
| 1 | GPH67635-RA | XM_018972085.1 | 甲基转移酶PMT26 Methyltransferase PMT26 |
| 2 | GPH79708-RA | DQ471302.1 | 可能的叶绿体叶绿素a/b结合蛋白 Putative chloroplast chlorophyll a/b-binding protein |
| 3 | GPH63128-RA | XM_018962628.1 | F-box蛋白 F-box protein SKIP27-like |
| 4 | GPH00558-RA | XM_018953092.1 | 木聚糖内转葡萄糖基酶/水解酶蛋白6 Xyloglucan endotransglucosylase/hydrolase protein 6 |
| 5 | GPH52393-RA | XM_018993881.1 | β-半乳糖苷酶 Beta-galactosidase |
| 6 | GPH12478-RA | XM_018999934.1 | L-抗坏血酸过氧化物酶6 L-ascorbate peroxidase 6 |
| 7 | GPH50419-RA | XM_018990252.1 | 甘氨酸脱氢酶 Glycine dehydrogenase |
| 8 | GPH19320-RA | XM_018980394.1 | 超敏诱导反应蛋白1 Hypersensitive-induced response protein 1-like |
| 9 | GPH69606-RA | XM_024066525.1 | 甲酸脱氢酶 Formate dehydrogenase |
| 10 | GPH62533-RA | XM_024031608.1 | 过氧化物酶42 Peroxidase 42 |
| 11 | GPH15474-RA | XM_018967583.1 | 肽基脯氨酰顺反异构酶 Peptidyl-prolyl cis-trans isomerase |
| 12 | GPH07814-RA | — | 光皮桦KNOX4转录因子 BlKNOX4 |
| 13 | GPH63909-RA | XM_018988520.1 | 丝氨酸羟甲基转移酶4 Serine hydroxymethyltransferase 4 |
| 14 | GPH42621-RA | XM_018967753.1 | 蛋白磷酸酶2C 23 Protein phosphatase 2C 23 |
| 15 | GPH08071-RA | XM_018969160.1 | 乙酰鸟氨酸脱乙酰酶 Acetylornithine deacetylase |
| 16 | GPH38087-RA | XM_018983680.1 | 组蛋白赖氨酸N-甲基转移酶2D Histone-lysine N-methyltransferase 2D |
| 17 | GPH33933-RA | XM_024050483.1 | DNA导向的RNA聚合酶Ⅱ DNA-directed RNA polymerases Ⅱ |
| 18 | GPH07814-RA | XM_024065073.1 | 同源异型盒knotted-1样蛋白LET6 Homeobox protein knotted-1-like LET6 |
| 19 | GPH77232-RA | KP711310.1 | 苯丙氨酸解氨酶2 Phenylalanine ammonia-lyase 2 (PAL2) |
| 20 | GPH25088-RA | XM_024029965.1 | 核糖核酸酶2 Ribonuclease 2 |
| 21 | GPH67318-RA | XM_018996369.1 | 细胞色素b6-f复合铁硫亚基1 Cytochrome b6-f complex iron-sulfur subunit 1 |
| 22 | GPH04217-RA | — | 光皮桦KNOX9转录因子 BlKNOX9 |
| 23 | GPH60229-RA | XM_018982373.1 | α, α-海藻糖磷酸合酶6 Alpha, alpha-trehalose-phosphate synthase 6 |
| 24 | GPH09215-RA | XM_024059006.1 | EXORDIUM样蛋白 EXORDIUM-like protein |
| 25 | GPH57276-RA | XM_018982529.1 | SAR1A样GTP结合蛋白 GTP-binding protein SAR1A-like |
| 26 | GPH08734-RA | XM_024042059.1 | rac样GTP结合蛋白5 Rac-like GTP-binding protein 5 |
| 27 | GPH77067-RA | XM_018990062.1 | WD-40重复序列蛋白 WD-40 repeat-containing protein MSI4-like |
| 28 | GPH39769-RA | XM_018953914.1 | 65kDa微管相关蛋白8 65-kDa microtubule-associated protein 8-like |
| 29 | GPH26685-RA | XM_024016523.1 | 核苷水解酶15 Nudix hydrolase 15 |
| 30 | GPH24400-RA | XM_018962128.1 | 40S核糖体蛋白 40S ribosomal protein S9-2-like |
| 31 | GPH75230-RA | — | 光皮桦STM转录因子 BlSTM |
| 32 | GPH10713-RA | XM_018988639.1 | 类糊精硫醇蛋白酶 Thiol protease aleurain-like |
| 33 | GPH72582-RA | XM_018991497.1 | 天冬酰胺-tRNA连接酶 Asparagine-tRNA ligase |
| 34 | GPH50474-RA | XM_018984817.1 | 酮酸还原异构酶 Ketol-acid reductoisomerase |
| 35 | GPH00799-RA | XM_024032479.1 | 真核翻译起始因子3 Eukaryotic translation initiation factor 3 |
| 36 | GPH41993-RA | XM_018966504.1 | 细胞色素P450 Cytochrome P450 84A1-like |
| 37 | GPH54999-RA | XM_024058733.1 | 钙调素 Calmodulin-7-like |
| 38 | GPH68179-RA | XM_019001046.1 | sm样蛋白LSM8 Sm-like protein LSM8 |
| 39 | GPH41167-RA | XM_024073889.1 | 脱水诱导样蛋白19 Protein DEHYDRATION-INDUCED 19-like |
| 40 | GPH04307-RA | XM_018957060.1 | SWI/SNF复合亚基SWI3B SWI/SNF complex subunit SWI3B |
| 41 | GPH64726-RA | XM_024063283.1 | LHCII型叶绿素a-b结合蛋白 Chlorophyll a-b binding protein of LHCII type 1-like |
| 42 | GPH19951-RA | XM_018964336.1 | 泛素蛋白连接酶E3 Ubiquitin-protein ligase E3 |
| 43 | GPH32771-RA | XM_024035128.1 | 40S核糖体蛋白 40S ribosomal protein S2-3-like |
| 44 | GPH42243-RA | XM_006449315.2 | 休眠相关蛋白1 Dormancy-associated protein 1 |
| 45 | GPH74993-RA | XM_018953684.1 | 外二元环裂解双加氧酶 Extradiol ring-cleavage dioxygenase-like |
| 46 | GPH43902-RA | XM_018973673.1 | 氨基酰化酶-1 Aminoacylase-1 |
| 47 | GPH20575-RA | XM_024047220.1 | GATA转录因子26 GATA transcription factor 26 |

Fig.5
Verification of interactions among BlBLH1 and three KNOX proteins by using BiFC (Bar=10 μm) The first column is yelow fluorescent signal of YFP in Arabidopsis protoplast cells, the second column is chloroplast spontaneous fluorescence, the third column is the bright-field image of Arabidopsis protoplast, and the fourth column is the merged image of YFP fluorescence, chloroplast spontaneous fluorescence and bright-field."


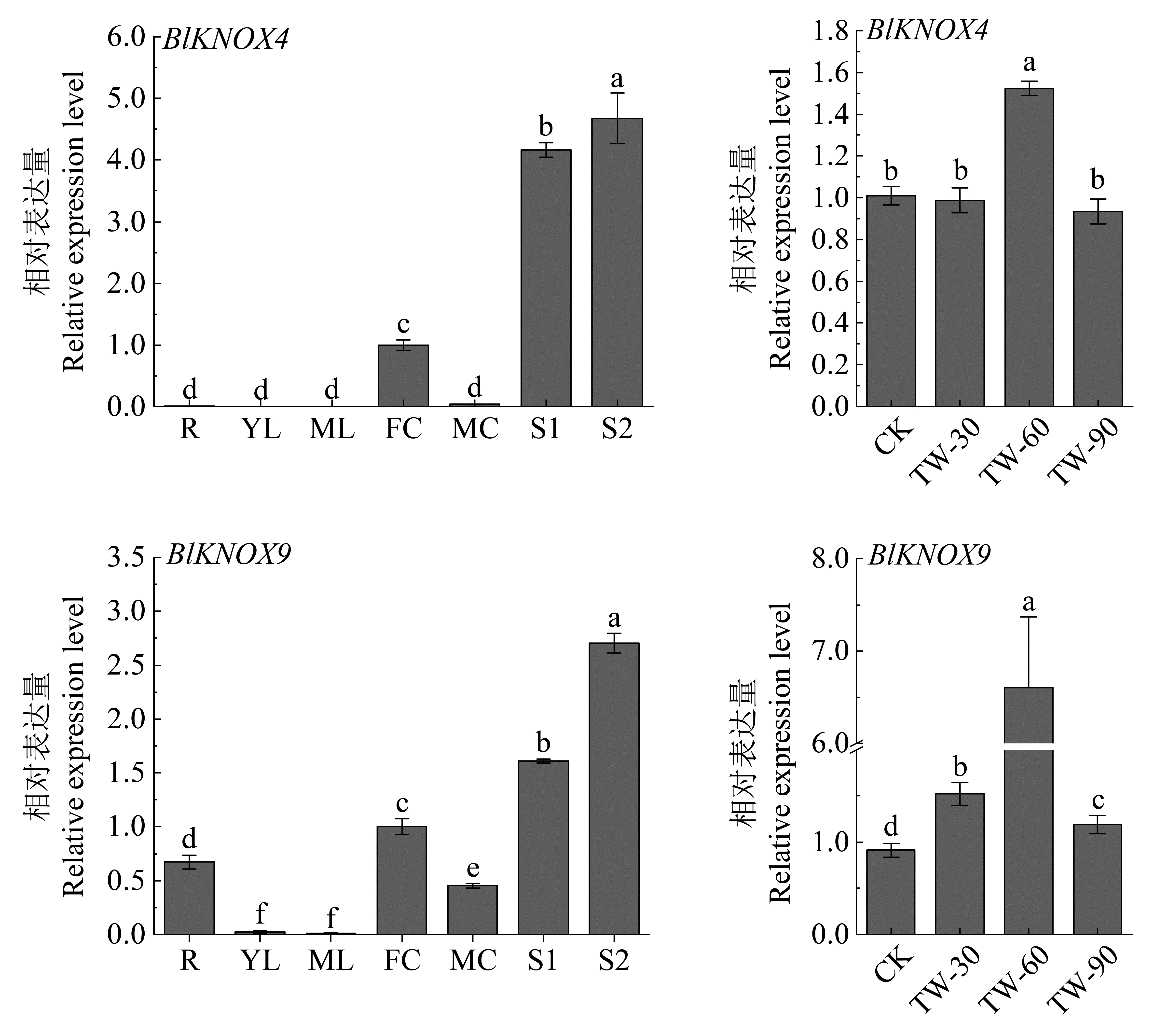
Fig.6
Expression analysis of BlKNOX4 and BlKNOX9 in different tissues, organs and treatment process of tension wood Different small letters above columns indicate significant difference at 0. 05 levels. R: Root; YL: Young leaf; ML: Mature leaf; FF: Female inflorescence; MF: Male inflorescence; S1: Non-lignified stem; S2: Lignified stem; TW: Tension wood. CK: Control."
| 赖叶林, 贺莹, 李欣欣, 等. 一种植物原生质体分离与瞬时转化的方法. 植物生理学报, 2020, 56 (4): 895- 903. | |
| Lai Y L , He Y , Li X X , et al. A method for isolation and transient transformation of plant protoplasts. Plant Physiology Journal, 2020, 56 (4): 895- 903. | |
| 刘文哲, 牛明月, 李秀云, 等. 光皮桦实时荧光定量PCR内参基因的筛选. 林业科学, 2016, 52 (8): 29- 37. | |
| Liu W Z , Niu M Y , Li X Y , et al. The selection of reference genes for quantitative PCR in Betula luminifera. Scientia Silvae Sinicae, 2016, 52 (8): 29- 37. | |
| 刘一星, 赵广杰. 木材学. 北京: 中国林业出版社, 2012. | |
| Liu Y X , Zhao G J . Wood Science. Beijing: China Forest Publishing House, 2012. | |
| 俞子承, 倪飞, 江成, 等. 光皮桦BlCCoAOMT基因的克隆, 表达及单核苷酸变异分析. 核农学报, 2019, 33 (5): 870- 879. | |
| Yu Z C , Ni F , Jiang C , et al. Cloning, expression and single nucleotide variation analysis of BlCCoAOMT gene in Betula luminifera. Journal of Nuclear Agricultural Sciences, 2019, 33 (5): 870- 879. | |
| 郑万钧. 中国树木志. 北京: 中国林业出版社, 1983. | |
| Zheng W J . Tree Index of China. Beijing: China Forest Publishing House, 1983. | |
|
Arnaud N , Pautot V . Ring the BELL and tie the KNOX: roles for TALEs in gynoecium development. Frontiers in Plant Science, 2014, 5 (93): 1- 7.
doi: 10.3389/fpls.2014.00093 |
|
|
Becker A , Bey M , Bürglin T R , et al. Ancestry and diversity of BEL1 -like homeobox genes revealed by gymnosperm (Gnetum gnemon) homologs. Development Genes and Evolution, 2002, 212 (9): 452- 457.
doi: 10.1007/s00427-002-0259-7 |
|
|
Bellaoui M , Pidkowich M S , Samach A , et al. The Arabidopsis BELL1 and KNOX TALE homeodomain proteins interact through a domain conserved between plants and animals. Plant Cell, 2001, 13 (11): 2455- 2470.
doi: 10.1105/tpc.010161 |
|
|
Bhatt A M , Etchells J P , Canales C , et al. VAAMANA—a BEL1-like homeodomain protein, interacts with KNOX proteins BP and STM and regulates inflorescence stem growth in Arabidopsis. Gene, 2004, 328 (1): 103- 111.
doi: 10.1016/j.gene.2003.12.033 |
|
| Cai M , Huang H , Ni F , et al. RNA-Seq analysis of differential gene expression in Betula luminifera xylem during the early stages of tension wood formation. PeerJ, 6 (1): 5427- 5454. | |
|
Chano V , Sobrino-Plata J , Collada C , et al. Wood development regulators involved in apical growth in Pinus canariensis. Plant Biology, 2021, 23 (3): 438- 444.
doi: 10.1111/plb.13228 |
|
|
Cole M , Nolte C , Werr W . Nuclear import of the transcription factor SHOOT MERISTEMLESS depends on heterodimerization with BLH proteins expressed in discrete sub-domains of the shoot apical meristem of Arabidopsis thaliana. Nucleic Acids Research, 2006, 34 (4): 1281- 1292.
doi: 10.1093/nar/gkl016 |
|
|
Hackbusch J , Richter K , Müller J , et al. A central role of Arabidopsis thaliana ovate family proteins in networking and subcellular localization of 3-aa loop extension homeodomain proteins. PNAS, 2005, 102 (13): 4908- 4912.
doi: 10.1073/pnas.0501181102 |
|
|
Hamant O , Pautot V . Plant development: a TALE story. Comptes Rendus Biologies, 2010, 333 (4): 371- 381.
doi: 10.1016/j.crvi.2010.01.015 |
|
|
Huang H , Jiang C , Tong Z , et al. Eight distinct cellulose synthase catalytic subunit genes from Betula luminifera are associated with primary and secondary cell wall biosynthesis. Cellulose, 2014, 21 (4): 2183- 2198.
doi: 10.1007/s10570-014-0261-z |
|
|
Kim D , Cho Y H , Ryu H , et al. BLH1 and KNAT3 modulate ABA responses during germination and early seedling development in Arabidopsis. Plant Journal, 2013, 75 (5): 755- 766.
doi: 10.1111/tpj.12236 |
|
| Liu Y . Functional analysis of homeodomain transcription factors in secondary cell wall formation in Arabidopsis thaliana. USA: University of British Columbia, 2015, | |
| Liu Y , Douglas C J . A role for ovate family protein1 (OFP1) and OFP4 in a BLH6-KNAT7 multi-protein complex regulating secondary cell wall formation in Arabidopsis thaliana. Plant Signaling & Behavior, 2015, 10 (7): 1559- 2324. | |
|
Liu Y , You S , Taylor-Teeples M , et al. BEL1-LIKE HOMEODOMAIN6 and KNOTTED ARABIDOPSIS THALIANA7 interact and regulate secondary cell wall formation via repression of REVOLUTA. Plant Cell, 2014, 26 (12): 4843- 4861.
doi: 10.1105/tpc.114.128322 |
|
|
Ma Q , Wang N , Hao P , et al. Genome-wide identification and characterization of TALE superfamily genes in cotton reveals their functions in regulating secondary cell wall biosynthesis. BMC Plant Biology, 2019, 19 (1): 432- 451.
doi: 10.1186/s12870-019-2026-1 |
|
|
Mukherjee K , Brocchieri L , Bürglin . A comprehensive classification and evolutionary analysis of plant homeobox genes. Molecular Biology and Evolution, 2009, 26 (12): 2775- 2794.
doi: 10.1093/molbev/msp201 |
|
| Plomion C , Leprovost G , Stokes A . Wood formation in trees. Plant Physiology, 2001, 127 (1): 1513- 1523. | |
|
Smith H M S , Hake S . The interaction of two homeobox genes, BREVIPEDICELLUS and PENNYWISE, regulates internode patterning in the Arabidopsis inflorescence. The Plant Cell, 2003, 15 (8): 1717- 1727.
doi: 10.1105/tpc.012856 |
|
|
Taylor-Teeples M , Lin L , deLucas M , et al. An Arabidopsis gene regulatory network for secondary cell wall synthesis. Nature, 2015, 517 (7536): 571- 575.
doi: 10.1038/nature14099 |
|
|
Yan C , Hu Z , Nie Z , et al. CcBLH6, a bell-like homeodomain-containing transcription factor, regulates the fruit lignification pattern. Planta, 2021, 253 (5): 90- 101.
doi: 10.1007/s00425-021-03610-7 |
|
|
Zhang L , Zhang X , Ju H , et al. Ovate family protein1 interaction with BLH3 regulates transition timing from vegetative to reproductive phase in Arabidopsis. Biochemical and Biophysical Research Communications, 2016, 470 (3): 492- 497.
doi: 10.1016/j.bbrc.2016.01.135 |
| [1] | Yuan Li,Jinhuan Chen,Zhao Jin,Jingya Hou,Yusong Jiang,Haitao Xing. Functions of NAC128 Gene from Populus trichocarpa in Secondary Cell Wall Formation [J]. Scientia Silvae Sinicae, 2020, 56(11): 62-72. |
| [2] | Wang Lei, Su Hongyan, Gu Liang, Chang Zhiyuan, Chen Na. Screening and Verification of PtMPKs Interacting with PtMKK4 of Populus trichocarpa [J]. Scientia Silvae Sinicae, 2015, 51(10): 60-66. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||